INAF, a protein required for transient receptor potential Ca(2+) channel function
- PMID: 10557345
- PMCID: PMC23972
- DOI: 10.1073/pnas.96.23.13474
INAF, a protein required for transient receptor potential Ca(2+) channel function
Abstract
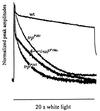
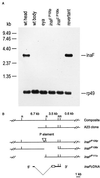
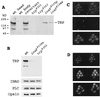
The trp gene of Drosophila encodes a subunit of a class of Ca(2+)-selective light-activated channels that carry the bulk of the phototransduction current. Transient receptor potential (TRP) homologs have been identified throughout animal phylogeny. In vertebrates, TRP-related channels have been suggested to mediate "store-operated Ca(2+) entry," which is important in Ca(2+) homeostasis in a wide variety of cell types. However, the mechanisms of activation and regulation of the TRP channel are not known. Here, we report on the Drosophila inaF gene, which encodes a highly eye-enriched protein, INAF, that appears to be required for TRP channel function. A null mutation in this gene significantly reduces the amount of the TRP protein and, in addition, specifically affects the TRP channel function so as to nearly shut down its activity. The inaF mutation also dramatically suppresses the severe degeneration caused by a constitutively active mutation in the trp gene. Although the reduction in the amount of the TRP protein may contribute to these phenotypes, several lines of evidence support the view that inaF mutations also more directly affect the TRP channel function, suggesting that the INAF protein may have a regulatory role in the channel function.
Figures


References
-
- Selinger Z, Minke B. Cold Spring Harbor Symp Quant Biol. 1988;53:333–341. - PubMed
-
- Bloomquist B B, Shortridge R D, Schneuwly S, Perdew M, Montell C, Steller H, Rubin G, Pak W L. Cell. 1988;54:723–733. - PubMed
-
- Toyoshima S, Matsumoto N, Wang P, Inoue H, Yoshioka T, Hotta Y, Osawa T. J Biol Chem. 1990;265:14842–14848. - PubMed
-
- Niemeyer B A, Suzuki E, Scott K, Jalink K, Zuker C S. Cell. 1996;85:651–659. - PubMed
-
- Hardie R C, Minke B. Neuron. 1992;8:613–651. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

