Arabidopsis thaliana PAD4 encodes a lipase-like gene that is important for salicylic acid signaling
- PMID: 10557364
- PMCID: PMC23991
- DOI: 10.1073/pnas.96.23.13583
Arabidopsis thaliana PAD4 encodes a lipase-like gene that is important for salicylic acid signaling
Abstract
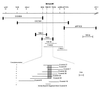
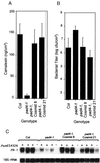
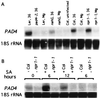
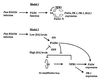
The Arabidopsis PAD4 gene previously was found to be required for expression of multiple defense responses including camalexin synthesis and PR-1 gene expression in response to infection by the bacterial pathogen Pseudomonas syringae pv. maculicola. This report describes the isolation of PAD4. The predicted PAD4 protein sequence displays similarity to triacyl glycerol lipases and other esterases. The PAD4 transcript was found to accumulate after P. syringae infection or treatment with salicylic acid (SA). PAD4 transcript levels were very low in infected pad4 mutants. Treatment with SA induced expression of PAD4 mRNA in pad4-1, pad4-3, and pad4-4 plants but not in pad4-2 plants. Induction of PAD4 expression by P. syringae was independent of the regulatory factor NPR1 but induction by SA was NPR1-dependent. Taken together with the previous observation that pad4 mutants have a defect in accumulation of SA upon pathogen infection, these results suggest that PAD4 participates in a positive regulatory loop that increases SA levels, thereby activating SA-dependent defense responses.
Figures

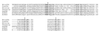

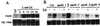
Similar articles
-
GBF1 differentially regulates CAT2 and PAD4 transcription to promote pathogen defense in Arabidopsis thaliana.Plant J. 2017 Sep;91(5):802-815. doi: 10.1111/tpj.13608. Epub 2017 Jul 10. Plant J. 2017. PMID: 28622438
-
A gain-of-function mutation in a plant disease resistance gene leads to constitutive activation of downstream signal transduction pathways in suppressor of npr1-1, constitutive 1.Plant Cell. 2003 Nov;15(11):2636-46. doi: 10.1105/tpc.015842. Epub 2003 Oct 23. Plant Cell. 2003. PMID: 14576290 Free PMC article.
-
Constitutive salicylic acid-dependent signaling in cpr1 and cpr6 mutants requires PAD4.Plant J. 2001 May;26(4):395-407. doi: 10.1046/j.1365-313x.2001.2641040.x. Plant J. 2001. PMID: 11439127
-
Genetic analysis of plant disease resistance pathways.Novartis Found Symp. 2001;236:153-61; discussion 161-4. doi: 10.1002/9780470515778.ch11. Novartis Found Symp. 2001. PMID: 11387977 Review.
-
Nitric oxide and salicylic acid signaling in plant defense.Proc Natl Acad Sci U S A. 2000 Aug 1;97(16):8849-55. doi: 10.1073/pnas.97.16.8849. Proc Natl Acad Sci U S A. 2000. PMID: 10922045 Free PMC article. Review.
Cited by
-
Multidimensional gene regulatory landscape of a bacterial pathogen in plants.Nat Plants. 2020 Jul;6(7):883-896. doi: 10.1038/s41477-020-0690-7. Epub 2020 Jun 15. Nat Plants. 2020. PMID: 32541952
-
Identification, characterization and mapping of differentially expressed genes in a winter wheat cultivar (Centenaire) resistant to Fusarium graminearum infection.Mol Biol Rep. 2012 Oct;39(10):9583-600. doi: 10.1007/s11033-012-1823-5. Epub 2012 Jun 21. Mol Biol Rep. 2012. PMID: 22718510
-
In silico study of interaction between rice proteins enhanced disease susceptibility 1 and phytoalexin deficient 4, the regulators of salicylic acid signalling pathway.J Biosci. 2012 Jul;37(3):563-71. doi: 10.1007/s12038-012-9208-4. J Biosci. 2012. PMID: 22750992
-
Fighting Asian Soybean Rust.Front Plant Sci. 2016 Jun 7;7:797. doi: 10.3389/fpls.2016.00797. eCollection 2016. Front Plant Sci. 2016. PMID: 27375652 Free PMC article. Review.
-
Early senescence and cell death in Arabidopsis saul1 mutants involves the PAD4-dependent salicylic acid pathway.Plant Physiol. 2012 Aug;159(4):1477-87. doi: 10.1104/pp.112.196220. Epub 2012 Jun 15. Plant Physiol. 2012. PMID: 22706448 Free PMC article.
References
-
- Baker B, Zambryski P, Staskawicz B, Dinesh-Kumar S P. Science. 1997;276:726–733. - PubMed
-
- Gaffney T, Friedrich L, Vernooij B, Negrotto D, Nye G, Uknes S, Ward E, Kessmann H, Ryals J. Science. 1993;261:754–756. - PubMed
-
- Delaney T P, Uknes S, Vernooij B, Friedrich L, Weymann K, Negrotto D, Gaffney T, Gut-Rella M, Kessman H, Ward E, et al. Science. 1994;266:1247–1250. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

