The activation domain of herpesvirus saimiri R protein interacts with the TATA-binding protein
- PMID: 10559285
- PMCID: PMC113022
- DOI: 10.1128/JVI.73.12.9756-9763.1999
The activation domain of herpesvirus saimiri R protein interacts with the TATA-binding protein
Abstract
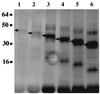
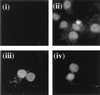
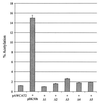
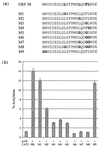
The herpesvirus saimiri open reading frame (ORF) 50 produces two transcripts. The first is spliced, contains a single intron, and is detected at early times during the productive cycle, whereas the second is expressed later and is produced from a promoter within the second exon. Analysis of their gene products has shown that they function as sequence specific transactivators. In this report, we demonstrate that the carboxy terminus of ORF 50b contains an activation domain which is essential for transactivation. This domain contains positionally conserved hydrophobic residues found in a number of activation domains, including the herpes simplex virus VP16 and the Epstein-Barr virus R proteins. Mutational analysis of this domain demonstrates that these conserved hydrophobic residues are essential for ORF 50 transactivation capability. Furthermore, this domain is required for the interaction between the ORF 50 proteins and the basal transcription factor TATA-binding protein.
Figures





Similar articles
-
The carboxy terminus of the herpesvirus saimiri ORF 57 gene contains domains that are required for transactivation and transrepression.J Gen Virol. 2000 Sep;81(Pt 9):2253-2265. doi: 10.1099/0022-1317-81-9-2253. J Gen Virol. 2000. PMID: 10950983
-
The herpesvirus saimiri ORF50 gene, encoding a transcriptional activator homologous to the Epstein-Barr virus R protein, is transcribed from two distinct promoters of different temporal phases.J Virol. 1997 Mar;71(3):2550-4. doi: 10.1128/JVI.71.3.2550-2554.1997. J Virol. 1997. PMID: 9032399 Free PMC article.
-
The Herpesvirus saimiri replication and transcription activator acts synergistically with CCAAT enhancer binding protein alpha to activate the DNA polymerase promoter.J Virol. 2005 Nov;79(21):13548-60. doi: 10.1128/JVI.79.21.13548-13560.2005. J Virol. 2005. PMID: 16227275 Free PMC article.
-
Open reading frame 50 protein of Kaposi's sarcoma-associated herpesvirus directly activates the viral PAN and K12 genes by binding to related response elements.J Virol. 2002 Apr;76(7):3168-78. doi: 10.1128/jvi.76.7.3168-3178.2002. J Virol. 2002. PMID: 11884541 Free PMC article.
-
The Epstein-Barr virus R transactivator (Rta) contains a complex, potent activation domain with properties different from those of VP16.J Virol. 1992 Sep;66(9):5500-8. doi: 10.1128/JVI.66.9.5500-5508.1992. J Virol. 1992. PMID: 1323708 Free PMC article.
Cited by
-
m6A Regulates the Stability of Cellular Transcripts Required for Efficient KSHV Lytic Replication.Viruses. 2023 Jun 16;15(6):1381. doi: 10.3390/v15061381. Viruses. 2023. PMID: 37376680 Free PMC article.
-
The latency-associated nuclear antigen homolog of herpesvirus saimiri inhibits lytic virus replication.J Virol. 2003 May;77(10):5911-25. doi: 10.1128/jvi.77.10.5911-5925.2003. J Virol. 2003. PMID: 12719584 Free PMC article.
-
The DNA architectural protein HMGB1 facilitates RTA-mediated viral gene expression in gamma-2 herpesviruses.J Virol. 2004 Dec;78(23):12940-50. doi: 10.1128/JVI.78.23.12940-12950.2004. J Virol. 2004. PMID: 15542646 Free PMC article.
-
Kaposi's sarcoma-associated herpesvirus RTA promotes degradation of the Hey1 repressor protein through the ubiquitin proteasome pathway.J Virol. 2009 Jul;83(13):6727-38. doi: 10.1128/JVI.00351-09. Epub 2009 Apr 15. J Virol. 2009. PMID: 19369342 Free PMC article.
-
Merkel Cell Polyomavirus Small T Antigen Drives Cell Motility via Rho-GTPase-Induced Filopodium Formation.J Virol. 2018 Jan 2;92(2):e00940-17. doi: 10.1128/JVI.00940-17. Print 2018 Jan 15. J Virol. 2018. PMID: 29093086 Free PMC article.
References
-
- Albrecht J C, Fleckenstein B. Structural organization of the conserved gene block of herpesvirus saimiri coding for DNA polymerase, glycoprotein B, and major DNA binding protein. Virology. 1990;174:533–542. - PubMed
-
- Bublot M, Manet E, Lequarre A S, Albrecht J C, Nicholas J, Fleckenstein B, Pastoret P P, Thiry E. Genetic relationships between bovine herpesvirus 4 and the gamma-herpesviruses Epstein-Barr and herpesvirus saimiri. Virology. 1992;190:654–665. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

