Translating ribosomes inhibit poliovirus negative-strand RNA synthesis
- PMID: 10559325
- PMCID: PMC113062
- DOI: 10.1128/JVI.73.12.10104-10112.1999
Translating ribosomes inhibit poliovirus negative-strand RNA synthesis
Abstract
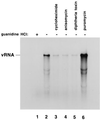
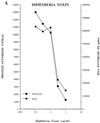
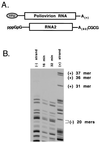
Poliovirus has a single-stranded RNA genome of positive polarity that serves two essential functions at the start of the viral replication cycle in infected cells. First, it is translated to synthesize viral proteins and, second, it is copied by the viral polymerase to synthesize negative-strand RNA. We investigated these two reactions by using HeLa S10 in vitro translation-RNA replication reactions. Preinitiation RNA replication complexes were isolated from these reactions and then used to measure the sequential synthesis of negative- and positive-strand RNAs in the presence of different protein synthesis inhibitors. Puromycin was found to stimulate RNA replication overall. In contrast, RNA replication was inhibited by diphtheria toxin, cycloheximide, anisomycin, and ricin A chain. Dose-response experiments showed that precisely the same concentration of a specific drug was required to inhibit protein synthesis and to either stimulate or inhibit RNA replication. This suggested that the ability of these drugs to affect RNA replication was linked to their ability to alter the normal clearance of translating ribosomes from the input viral RNA. Consistent with this idea was the finding that the protein synthesis inhibitors had no measurable effect on positive-strand synthesis in normal RNA replication complexes. In marked contrast, negative-strand synthesis was stimulated by puromycin and was inhibited by cycloheximide. Puromycin causes polypeptide chain termination and induces the dissociation of polyribosomes from mRNA. Cycloheximide and other inhibitors of polypeptide chain elongation "freeze" ribosomes on mRNA and prevent the normal clearance of ribosomes from viral RNA templates. Therefore, it appears that the poliovirus polymerase was not able to dislodge translating ribosomes from viral RNA templates and mediate the switch from translation to negative-strand synthesis. Instead, the initiation of negative-strand synthesis appears to be coordinately regulated with the natural clearance of translating ribosomes to avoid the dilemma of ribosome-polymerase collisions.
Figures





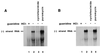
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

