Restricted immunoglobulin variable region (Ig V) gene expression accompanies secondary rearrangements of light chain Ig V genes in mouse plasmacytomas
- PMID: 10562316
- PMCID: PMC2195694
- DOI: 10.1084/jem.190.10.1405
Restricted immunoglobulin variable region (Ig V) gene expression accompanies secondary rearrangements of light chain Ig V genes in mouse plasmacytomas
Abstract
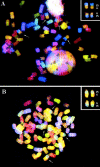
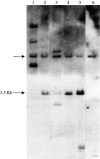
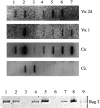
The many binding studies of monoclonal immunoglobulin (Ig) produced by plasmacytomas have found no universally common binding properties, but instead, groups of plasmacytomas with specific antigen-binding activities to haptens such as phosphorylcholine, dextrans, fructofuranans, or dinitrophenyl. Subsequently, it was found that plasmacytomas with similar binding chain specificities not only expressed the same idiotype, but rearranged the same light (V(L)) and heavy (V(H)) variable region genes to express a characteristic monoclonal antibody. In this study, we have examined by enzyme-linked immunosorbent assay five antibodies secreted by silicone-induced mouse plasmacytomas using a broader panel of antigens including actin, myosin, tubulin, single-stranded DNA, and double-stranded DNA. We have determined the Ig heavy and light chain V gene usage in these same plasmacytomas at the DNA and RNA level. Our studies reveal: (a) antibodies secreted by plasmacytomas bind to different antigens in a manner similar to that observed for natural autoantibodies; (b) the expressed Ig heavy genes are restricted in V gene usage to the V(H)-J558 family; and (c) secondary rearrangements occur at the light chain level with at least three plasmacytomas expressing both kappa and lambda light chain genes. These results suggest that plasmacytomas use a restricted population of B cells that may still be undergoing rearrangement, thereby bypassing the allelic exclusion normally associated with expression of antibody genes.
Figures


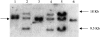



References
-
- Potter M., Wiener F. Plasmacytomagenesis in micemodel of neoplastic development dependent upon chromosomal translocations. Carcinogenesis. 1992;13:1681–1697. - PubMed
-
- Potter M. Antigen-binding myeloma proteins of mice. Adv. Immunol. 1977;25:141–211. - PubMed
-
- Guilbert B., Dighiero G., Avrameas S. Naturally occurring antibodies against nine common antigens in human sera. I. Detection, isolation and characterization. J. Immunol. 1982;128:2779–2787. - PubMed
-
- Dighiero G., Lymberi P., Mazie J.C., Rouyre S., Butler B.G., Whalen R.G., Avrameas S. Murine hybridomas secreting natural monoclonal antibodies reacting with self antigens. J. Immunol. 1983;131:2267–2272. - PubMed
-
- Diaw L., Magnac C., Pritsch O., Buckle M., Alzari P.M., Dighiero G. Structural and affinity studies of IgM polyreactive natural autoantibodies. J. Immunol. 1997;158:968–976. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

