The ATPase domain but not the acidic region of Cockayne syndrome group B gene product is essential for DNA repair
- PMID: 10564257
- PMCID: PMC25641
- DOI: 10.1091/mbc.10.11.3583
The ATPase domain but not the acidic region of Cockayne syndrome group B gene product is essential for DNA repair
Abstract
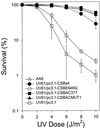
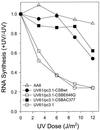
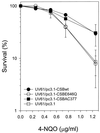
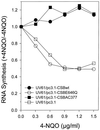
Cockayne syndrome (CS) is a human genetic disorder characterized by UV sensitivity, developmental abnormalities, and premature aging. Two of the genes involved, CSA and CSB, are required for transcription-coupled repair (TCR), a subpathway of nucleotide excision repair that removes certain lesions rapidly and efficiently from the transcribed strand of active genes. CS proteins have also been implicated in the recovery of transcription after certain types of DNA damage such as those lesions induced by UV light. In this study, site-directed mutations have been introduced to the human CSB gene to investigate the functional significance of the conserved ATPase domain and of a highly acidic region of the protein. The CSB mutant alleles were tested for genetic complementation of UV-sensitive phenotypes in the human CS-B homologue of hamster UV61. In addition, the CSB mutant alleles were tested for their ability to complement the sensitivity of UV61 cells to the carcinogen 4-nitroquinoline-1-oxide (4-NQO), which introduces bulky DNA adducts repaired by global genome repair. Point mutation of a highly conserved glutamic acid residue in ATPase motif II abolished the ability of CSB protein to complement the UV-sensitive phenotypes of survival, RNA synthesis recovery, and gene-specific repair. These data indicate that the integrity of the ATPase domain is critical for CSB function in vivo. Likewise, the CSB ATPase point mutant failed to confer cellular resistance to 4-NQO, suggesting that ATP hydrolysis is required for CSB function in a TCR-independent pathway. On the contrary, a large deletion of the acidic region of CSB protein did not impair the genetic function in the processing of either UV- or 4-NQO-induced DNA damage. Thus the acidic region of CSB is likely to be dispensable for DNA repair, whereas the ATPase domain is essential for CSB function in both TCR-dependent and -independent pathways.
Figures



Similar articles
-
Cockayne syndrome group B cellular and biochemical functions.Am J Hum Genet. 2003 Dec;73(6):1217-39. doi: 10.1086/380399. Epub 2003 Nov 24. Am J Hum Genet. 2003. PMID: 14639525 Free PMC article. Review.
-
Role of the ATPase domain of the Cockayne syndrome group B protein in UV induced apoptosis.Oncogene. 2000 Jan 27;19(4):477-89. doi: 10.1038/sj.onc.1203372. Oncogene. 2000. PMID: 10698517
-
Differential requirement for the ATPase domain of the Cockayne syndrome group B gene in the processing of UV-induced DNA damage and 8-oxoguanine lesions in human cells.Nucleic Acids Res. 2002 Feb 1;30(3):782-93. doi: 10.1093/nar/30.3.782. Nucleic Acids Res. 2002. PMID: 11809892 Free PMC article.
-
Phenotypic consequences of mutations in the conserved motifs of the putative helicase domain of the human Cockayne syndrome group B gene.Gene. 2002 Jan 23;283(1-2):27-40. doi: 10.1016/s0378-1119(01)00870-8. Gene. 2002. PMID: 11867210
-
The role of Cockayne Syndrome group B (CSB) protein in base excision repair and aging.Mech Ageing Dev. 2008 Jul-Aug;129(7-8):441-8. doi: 10.1016/j.mad.2008.04.009. Epub 2008 Apr 30. Mech Ageing Dev. 2008. PMID: 18541289 Free PMC article. Review.
Cited by
-
Molecular characterization of an acidic region deletion mutant of Cockayne syndrome group B protein.Nucleic Acids Res. 2000 Aug 15;28(16):3151-9. doi: 10.1093/nar/28.16.3151. Nucleic Acids Res. 2000. PMID: 10931931 Free PMC article.
-
Tethering of the conserved piggyBac transposase fusion protein CSB-PGBD3 to chromosomal AP-1 proteins regulates expression of nearby genes in humans.PLoS Genet. 2012 Sep;8(9):e1002972. doi: 10.1371/journal.pgen.1002972. Epub 2012 Sep 27. PLoS Genet. 2012. PMID: 23028371 Free PMC article.
-
Elements That Regulate the DNA Damage Response of Proteins Defective in Cockayne Syndrome.J Mol Biol. 2016 Jan 16;428(1):62-78. doi: 10.1016/j.jmb.2015.11.020. Epub 2015 Nov 23. J Mol Biol. 2016. PMID: 26616585 Free PMC article.
-
Cockayne syndrome group B cellular and biochemical functions.Am J Hum Genet. 2003 Dec;73(6):1217-39. doi: 10.1086/380399. Epub 2003 Nov 24. Am J Hum Genet. 2003. PMID: 14639525 Free PMC article. Review.
-
Cockayne syndrome B protein stimulates apurinic endonuclease 1 activity and protects against agents that introduce base excision repair intermediates.Nucleic Acids Res. 2007;35(12):4103-13. doi: 10.1093/nar/gkm404. Epub 2007 Jun 12. Nucleic Acids Res. 2007. PMID: 17567611 Free PMC article.
References
-
- Bohr VA, Okumoto DS. Analysis of pyrmidine dimers in defined genes. In: Friedberg EC, Hanawalt PC, editors. A Laboratory Manual of Research Procedures. New York: Marcel Dekker; 1988. pp. 347–366.
-
- Bohr VA, Smith CA, Okumoto DS, Hanawalt PC. DNA repair in an active gene: removal of pyrimidine dimers from the DHFR gene of CHO cells is much more efficient than in the genome overall. Cell. 1985;40:359–369. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

