Orientation of the transcription preinitiation complex in archaea
- PMID: 10570129
- PMCID: PMC24121
- DOI: 10.1073/pnas.96.24.13662
Orientation of the transcription preinitiation complex in archaea
Abstract
The basal transcription machinery of Archaea corresponds to the minimal subset of factors required for RNA polymerase II transcription in eukaryotes. Using just two factors, Archaea recruit the RNA polymerase to promoters and define the direction of transcription. Notably, the principal determinant for the orientation of transcription is not the recognition of the TATA box by the TATA-box-binding protein. Instead, transcriptional polarity is governed by the interaction of the archaeal TFIIB homologue with a conserved motif immediately upstream of the TATA box. This interaction yields an archaeal preinitiation complex with the same orientation as the analogous eukaryal complex.
Figures


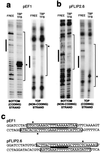


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

