In silico detection of control signals: mRNA 3'-end-processing sequences in diverse species
- PMID: 10570197
- PMCID: PMC24189
- DOI: 10.1073/pnas.96.24.14055
In silico detection of control signals: mRNA 3'-end-processing sequences in diverse species
Abstract
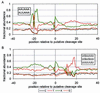
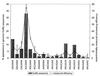
We have investigated mRNA 3'-end-processing signals in each of six eukaryotic species (yeast, rice, arabidopsis, fruitfly, mouse, and human) through the analysis of more than 20,000 3'-expressed sequence tags. The use and conservation of the canonical AAUAAA element vary widely among the six species and are especially weak in plants and yeast. Even in the animal species, the AAUAAA signal does not appear to be as universal as indicated by previous studies. The abundance of single-base variants of AAUAAA correlates with their measured processing efficiencies. As found previously, the plant polyadenylation signals are more similar to those of yeast than to those of animals, with both common content and arrangement of the signal elements. In all species examined, the complete polyadenylation signal appears to consist of an aggregate of multiple elements. In light of these and previous results, we present a broadened concept of 3'-end-processing signals in which no single exact sequence element is universally required for processing. Rather, the total efficiency is a function of all elements and, importantly, an inefficient word in one element can be compensated for by strong words in other elements. These complex patterns indicate that effective tools to identify 3'-end-processing signals will require more than consensus sequence identification.
Figures


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

