Receptor-mediated endocytosis in the Caenorhabditis elegans oocyte
- PMID: 10588660
- PMCID: PMC25760
- DOI: 10.1091/mbc.10.12.4311
Receptor-mediated endocytosis in the Caenorhabditis elegans oocyte
Abstract
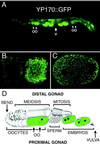
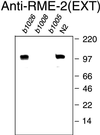
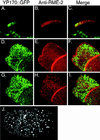
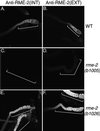
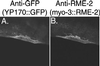
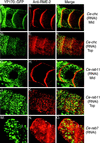
The Caenorhabditis elegans oocyte is a highly amenable system for forward and reverse genetic analysis of receptor-mediated endocytosis. We describe the use of transgenic strains expressing a vitellogenin::green fluorescent protein (YP170::GFP) fusion to monitor yolk endocytosis by the C. elegans oocyte in vivo. This YP170::GFP reporter was used to assay the functions of C. elegans predicted proteins homologous to vertebrate endocytosis factors using RNA-mediated interference. We show that the basic components and pathways of endocytic trafficking are conserved between C. elegans and vertebrates, and that this system can be used to test the endocytic functions of any new gene. We also used the YP170::GFP assay to identify rme (receptor-mediated endocytosis) mutants. We describe a new member of the low-density lipoprotein receptor superfamily, RME-2, identified in our screens for endocytosis defective mutants. We show that RME-2 is the C. elegans yolk receptor.
Figures



Comment in
-
An MBoC favorite: receptor-mediated endocytosis in the Caenorhabditis elegans oocyte.Mol Biol Cell. 2012 Jun;23(12):2235. doi: 10.1091/mbc.E12-02-0143. Mol Biol Cell. 2012. PMID: 22695480 Free PMC article. No abstract available.
References
-
- Bettinger JC, Lee K, Rougvie AE. Stage-specific accumulation of the terminal differentiation factor LIN-29 during Caenorhabditis elegans development. Development. 1996;122:2517–2527. - PubMed
-
- Bossinger O, Schierenberg E. Cell-cell communication in the embryo of Caenorhabditis elegans. Dev Biol. 1992;151:401–409. - PubMed
-
- Bossinger O, Schierenberg E. The use of fluorescent marker dyes for studying intercellular communication in nematode embryos. Int J Dev Biol. 1996;40:431–439. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

