Specific mutations in a viral RNA pseudoknot drastically change ribosomal frameshifting efficiency
- PMID: 10588689
- PMCID: PMC24420
- DOI: 10.1073/pnas.96.25.14234
Specific mutations in a viral RNA pseudoknot drastically change ribosomal frameshifting efficiency
Abstract
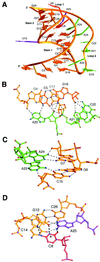
Many viruses regulate protein synthesis by -1 ribosomal frameshifting using an RNA pseudoknot. Frameshifting is vital for viral reproduction. Using the information gained from the recent high-resolution crystal structure of the beet western yellow virus pseudoknot, a systematic mutational analysis has been carried out in vitro and in vivo. We find that specific nucleotide tertiary interactions at the junction between the two stems of the pseudoknot are crucial. A triplex is found between stem 1 and loop 2, and triplex interactions are required for frameshifting function. For some mutations, loss of one hydrogen bond is sufficient to abolish frameshifting. Furthermore, mutations near the 5' end of the pseudoknot can increase frameshifting by nearly 300%, possibly by modifying ribosomal contacts. It is likely that the selection of suitable mutations can thus allow viruses to adjust frameshifting efficiencies and thereby regulate protein synthesis in response to environmental change.
Figures


Comment in
-
Programmed ribosomal frameshifting: much ado about knotting!Proc Natl Acad Sci U S A. 1999 Dec 7;96(25):14177-9. doi: 10.1073/pnas.96.25.14177. Proc Natl Acad Sci U S A. 1999. PMID: 10588670 Free PMC article. No abstract available.
Similar articles
-
Prokaryotic and eukaryotic translational machineries respond differently to the frameshifting RNA signal from plant or animal virus.Virus Res. 2003 Apr;92(2):165-70. doi: 10.1016/s0168-1702(03)00042-x. Virus Res. 2003. PMID: 12686425 Free PMC article. Review.
-
Minor groove RNA triplex in the crystal structure of a ribosomal frameshifting viral pseudoknot.Nat Struct Biol. 1999 Mar;6(3):285-92. doi: 10.1038/6722. Nat Struct Biol. 1999. PMID: 10074948 Free PMC article.
-
Structural and functional studies of retroviral RNA pseudoknots involved in ribosomal frameshifting: nucleotides at the junction of the two stems are important for efficient ribosomal frameshifting.EMBO J. 1995 Feb 15;14(4):842-52. doi: 10.1002/j.1460-2075.1995.tb07062.x. EMBO J. 1995. PMID: 7882986 Free PMC article.
-
Mutational analysis of the RNA pseudoknot component of a coronavirus ribosomal frameshifting signal.J Mol Biol. 1991 Aug 20;220(4):889-902. doi: 10.1016/0022-2836(91)90361-9. J Mol Biol. 1991. PMID: 1880803 Free PMC article.
-
A review on architecture of the gag-pol ribosomal frameshifting RNA in human immunodeficiency virus: a variability survey of virus genotypes.J Biomol Struct Dyn. 2017 Jun;35(8):1629-1653. doi: 10.1080/07391102.2016.1194231. Epub 2016 Aug 2. J Biomol Struct Dyn. 2017. PMID: 27485859 Review.
Cited by
-
A genome-wide analysis of RNA pseudoknots that stimulate efficient -1 ribosomal frameshifting or readthrough in animal viruses.Biomed Res Int. 2013;2013:984028. doi: 10.1155/2013/984028. Epub 2013 Nov 4. Biomed Res Int. 2013. PMID: 24298557 Free PMC article.
-
The global structures of a wild-type and poorly functional plant luteoviral mRNA pseudoknot are essentially identical.RNA. 2006 Nov;12(11):1959-69. doi: 10.1261/rna.199006. Epub 2006 Sep 25. RNA. 2006. PMID: 17000902 Free PMC article.
-
A loop 2 cytidine-stem 1 minor groove interaction as a positive determinant for pseudoknot-stimulated -1 ribosomal frameshifting.Proc Natl Acad Sci U S A. 2005 Sep 6;102(36):12694-9. doi: 10.1073/pnas.0506166102. Epub 2005 Aug 25. Proc Natl Acad Sci U S A. 2005. PMID: 16123125 Free PMC article.
-
An intermolecular RNA triplex provides insight into structural determinants for the pseudoknot stimulator of -1 ribosomal frameshifting.Nucleic Acids Res. 2010 Mar;38(5):1676-85. doi: 10.1093/nar/gkp1107. Epub 2009 Dec 8. Nucleic Acids Res. 2010. PMID: 20007152 Free PMC article.
-
Prokaryotic and eukaryotic translational machineries respond differently to the frameshifting RNA signal from plant or animal virus.Virus Res. 2003 Apr;92(2):165-70. doi: 10.1016/s0168-1702(03)00042-x. Virus Res. 2003. PMID: 12686425 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

