Further examination of the Xist promoter-switch hypothesis in X inactivation: evidence against the existence and function of a P(0) promoter
- PMID: 10588721
- PMCID: PMC24452
- DOI: 10.1073/pnas.96.25.14424
Further examination of the Xist promoter-switch hypothesis in X inactivation: evidence against the existence and function of a P(0) promoter
Abstract
The onset of X inactivation coincides with accumulation of Xist RNA along the future inactive X chromosome. A recent hypothesis proposed that accumulation is initiated by a promoter switch within Xist. In this hypothesis, an upstream promoter (P(0)) produces an unstable transcript, while the known downstream promoter (P(1)) produces a stable RNA. To test this hypothesis, we examined expression and half-life of Xist RNA produced from an Xist transgene lacking P(0) but retaining P(1). We confirm the previous finding that P(0) is dispensable for Xist expression in undifferentiated cells and that P(1) can be used in both undifferentiated and differentiated cells. Herein, we show that Xist RNA initiated at P(1) is unstable and does not accumulate. Further analysis indicates that the transcriptional boundary at P(0) does not represent the 5' end of a distinct Xist isoform. Instead, P(0) is an artifact of cross-amplification caused by a pseudogene of the highly expressed ribosomal protein S12 gene Rps12. Using strand-specific techniques, we find that transcription upstream of P(1) originates from the DNA strand opposite Xist and represents the 3' end of the antisense Tsix RNA. Thus, these data do not support the existence of a P(0) promoter and suggest that mechanisms other than switching of functionally distinct promoters control the up-regulation of Xist.
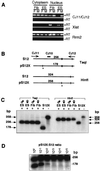
Figures




Similar articles
-
Developmentally regulated Xist promoter switch mediates initiation of X inactivation.Cell. 1998 Sep 18;94(6):809-17. doi: 10.1016/s0092-8674(00)81739-0. Cell. 1998. PMID: 9753327
-
Tsix-mediated repression of Xist accumulation is not sufficient for normal random X inactivation.Hum Mol Genet. 2001 Jun 15;10(13):1403-11. doi: 10.1093/hmg/10.13.1403. Hum Mol Genet. 2001. PMID: 11440993
-
A developmental switch in H4 acetylation upstream of Xist plays a role in X chromosome inactivation.EMBO J. 1999 May 17;18(10):2897-907. doi: 10.1093/emboj/18.10.2897. EMBO J. 1999. PMID: 10329635 Free PMC article.
-
Mammalian X chromosome inactivation.Novartis Found Symp. 1998;214:200-9; discussion 209-13, 228-32. doi: 10.1002/9780470515501.ch12. Novartis Found Symp. 1998. PMID: 9601019 Review.
-
X inactivation: Tsix and Xist as yin and yang.Curr Biol. 2000 Dec 14-28;10(24):R899-903. doi: 10.1016/s0960-9822(00)00847-2. Curr Biol. 2000. PMID: 11137025 Review.
Cited by
-
Characterization of the genomic Xist locus in rodents reveals conservation of overall gene structure and tandem repeats but rapid evolution of unique sequence.Genome Res. 2001 May;11(5):833-49. doi: 10.1101/gr.174901. Genome Res. 2001. PMID: 11337478 Free PMC article.
References
-
- Lyon M F. Nature (London) 1961;190:372–373. - PubMed
-
- Brown C J, Hendrich B D, Rupert J L, Lafreniere R G, Xing Y, Lawrence J, Willard H F. Cell. 1992;71:527–542. - PubMed
-
- Brockdorff N, Ashworth A, Kay G F, McCabe V M, Norris D P, Cooper P J, Swift S, Rastan S. Cell. 1992;71:515–526. - PubMed
-
- Penny G D, Kay G F, Sheardown S A, Rastan S, Brockdorff N. Nature (London) 1996;379:131–137. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

