Rsp5, a ubiquitin-protein ligase, is involved in degradation of the single-stranded-DNA binding protein rfa1 in Saccharomyces cerevisiae
- PMID: 10594025
- PMCID: PMC85078
- DOI: 10.1128/MCB.20.1.224-232.2000
Rsp5, a ubiquitin-protein ligase, is involved in degradation of the single-stranded-DNA binding protein rfa1 in Saccharomyces cerevisiae
Abstract
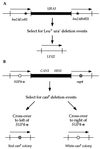
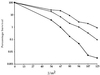
In Saccharomyces cerevisiae, RAD1 and RAD52 are required for alternate pathways of mitotic recombination. Double-mutant strains exhibit a synergistic interaction that decreases direct repeat recombination rates dramatically. A mutation in RFA1, the largest subunit of a single-stranded DNA-binding protein complex (RP-A), suppresses the recombination deficiency of rad1 rad52 strains (J. Smith and R. Rothstein, Mol. Cell. Biol. 15:1632-1641, 1995). Previously, we hypothesized that this mutation, rfa1-D228Y, causes an increase in recombinogenic lesions as well as the activation of a RAD52-independent recombination pathway. To identify gene(s) acting in this pathway, temperature-sensitive (ts) mutations were screened for those that decrease recombination levels in a rad1 rad52 rfa1-D228Y strain. Three mutants were isolated. Each segregates as a single recessive gene. Two are allelic to RSP5, which encodes an essential ubiquitin-protein ligase. One allele, rsp5-25, contains two mutations within its open reading frame. The first mutation does not alter the amino acid sequence of Rsp5, but it decreases the amount of full-length protein in vivo. The second mutation results in the substitution of a tryptophan with a leucine residue in the ubiquitination domain. In rsp5-25 mutants, the UV sensitivity of rfa1-D228Y is suppressed to the same level as in strains overexpressing Rfa1-D228Y. Measurement of the relative rate of protein turnover demonstrated that the half-life of Rfa1-D228Y in rsp5-25 mutants was extended to 65 min compared to a 35-min half-life in wild-type strains. We propose that Rsp5 is involved in the degradation of Rfa1 linking ubiquitination with the replication-recombination machinery.
Figures


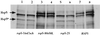


Similar articles
-
A mutation in the gene encoding the Saccharomyces cerevisiae single-stranded DNA-binding protein Rfa1 stimulates a RAD52-independent pathway for direct-repeat recombination.Mol Cell Biol. 1995 Mar;15(3):1632-41. doi: 10.1128/MCB.15.3.1632. Mol Cell Biol. 1995. PMID: 7862154 Free PMC article.
-
An allele of RFA1 suppresses RAD52-dependent double-strand break repair in Saccharomyces cerevisiae.Genetics. 1999 Feb;151(2):447-58. doi: 10.1093/genetics/151.2.447. Genetics. 1999. PMID: 9927442 Free PMC article.
-
A novel allele of Saccharomyces cerevisiae RFA1 that is deficient in recombination and repair and suppressible by RAD52.Mol Cell Biol. 1995 Mar;15(3):1620-31. doi: 10.1128/MCB.15.3.1620. Mol Cell Biol. 1995. PMID: 7862153 Free PMC article.
-
Studies of the interaction between Rad52 protein and the yeast single-stranded DNA binding protein RPA.Mol Cell Biol. 1998 Jul;18(7):4400-6. doi: 10.1128/MCB.18.7.4400. Mol Cell Biol. 1998. PMID: 9632824 Free PMC article.
-
Bul1, a new protein that binds to the Rsp5 ubiquitin ligase in Saccharomyces cerevisiae.Mol Cell Biol. 1996 Jul;16(7):3255-63. doi: 10.1128/MCB.16.7.3255. Mol Cell Biol. 1996. PMID: 8668140 Free PMC article.
Cited by
-
Sgs1 function in the repair of DNA replication intermediates is separable from its role in homologous recombinational repair.EMBO J. 2009 Apr 8;28(7):915-25. doi: 10.1038/emboj.2009.28. Epub 2009 Feb 12. EMBO J. 2009. PMID: 19214189 Free PMC article.
-
Localization of the Rsp5p ubiquitin-protein ligase at multiple sites within the endocytic pathway.Mol Cell Biol. 2001 May;21(10):3564-75. doi: 10.1128/MCB.21.10.3564-3575.2001. Mol Cell Biol. 2001. PMID: 11313482 Free PMC article.
-
A novel allele of fission yeast rad11 that causes defects in DNA repair and telomere length regulation.Nucleic Acids Res. 2003 Dec 15;31(24):7141-9. doi: 10.1093/nar/gkg917. Nucleic Acids Res. 2003. PMID: 14654689 Free PMC article.
-
SUMOylation regulates the homologous to E6-AP carboxyl terminus (HECT) ubiquitin ligase Rsp5p.J Biol Chem. 2013 Apr 12;288(15):10308-17. doi: 10.1074/jbc.M112.424234. Epub 2013 Feb 26. J Biol Chem. 2013. PMID: 23443663 Free PMC article.
-
The rad52-Y66A allele alters the choice of donor template during spontaneous chromosomal recombination.DNA Repair (Amst). 2010 Jan 2;9(1):23-32. doi: 10.1016/j.dnarep.2009.10.001. Epub 2009 Nov 4. DNA Repair (Amst). 2010. PMID: 19892607 Free PMC article.
References
-
- Aboussekhra A, Biggerstaff M, Shivji M K, Vilpo J A, Moncollin V, Podust V N, Protic M, Hubscher U, Egly J M, Wood R D. Mammalian DNA nucleotide excision repair reconstituted with purified protein components. Cell. 1995;80:859–868. - PubMed
-
- Benson F E, Baumann P, West S C. Synergistic actions of Rad51 and Rad52 in recombination and DNA repair. Nature. 1998;391:401–404. - PubMed
-
- Brill S J, Stillman B. Replication factor-A from Saccharomyces cerevisiae is encoded by three essential genes coordinately expressed at S phase. Genes Dev. 1991;5:1589–1600. - PubMed
-
- Brill S J, Stillman B. Yeast replication factor-A functions in the unwinding of the SV40 origin of DNA replication. Nature. 1989;342:92–95. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
