The dhfr oribeta-binding protein RIP60 contains 15 zinc fingers: DNA binding and looping by the central three fingers and an associated proline-rich region
- PMID: 10606657
- PMCID: PMC102514
- DOI: 10.1093/nar/28.2.570
The dhfr oribeta-binding protein RIP60 contains 15 zinc fingers: DNA binding and looping by the central three fingers and an associated proline-rich region
Abstract
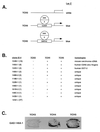
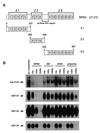
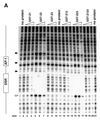
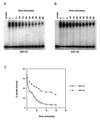
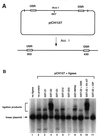
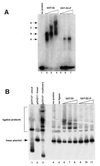
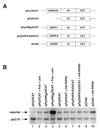
Initiation of DNA replication occurs with high frequency within oribeta, a short region 3' to the Chinese hamster dhfr gene. Homodimers of RIP60 (replication initiation-region protein 60 kDA) purified from nuclear extract bind two ATT-rich sites in oribeta and foster the formation of a twisted 720 bp DNA loop in vitro. Using a one hybrid screen in yeast, we have cloned the cDNA for human RIP60. RIP60 contains 15 C(2)H(2)zinc finger (ZF) DNA binding motifs organized in three clusters, termed hand Z1 (ZFs 1-5), hand Z2 (ZFs 6-8) and hand Z3 (ZFs 9-15). A proline-rich region is located between hands Z2 and Z3. Gel mobility shift and DNase I footprinting experiments show hands Z1 and Z2 independently bind the oribeta RIP60 sites specifically, but with different affinities. Hand Z3 binds DNA, but displays no specificity for RIP60 sites. Ligation enhancement, DNase I footprinting, and atomic force microscopy assays show that hand Z2 and a portion of the associated proline-rich region is sufficient for protein multimerization on DNA and DNA looping in vitro. Polyomavirus origin-dependent plasmid replication assays show RIP60 has weak replication enhancer activity, suggesting that RIP60 does not harbor a transcriptional transactivation domain. Because vertebrate origins of replication have no known consensus sequence, we suggest that sequence-specific DNA binding proteins such as RIP60 may act as accessory factors in origin identification prior to the assembly of pre-initiation complexes.
Figures



Similar articles
-
Condensation by DNA looping facilitates transfer of large DNA molecules into mammalian cells.Nucleic Acids Res. 2001 May 1;29(9):1982-8. doi: 10.1093/nar/29.9.1982. Nucleic Acids Res. 2001. PMID: 11328883 Free PMC article.
-
Chemical footprinting of structural and functional elements of dhfr oribeta during the CHOC 400 cell cycle.Gene. 2004 May 12;332:139-47. doi: 10.1016/j.gene.2004.02.032. Gene. 2004. PMID: 15145063
-
RIP60, a mammalian origin-binding protein, enhances DNA bending near the dihydrofolate reductase origin of replication.Mol Cell Biol. 1990 Dec;10(12):6236-43. doi: 10.1128/mcb.10.12.6236-6243.1990. Mol Cell Biol. 1990. PMID: 2247056 Free PMC article.
-
RIP60 dimers and multiples of dimers assemble link structures at an origin of bidirectional replication in the dihydrofolate reductase amplicon of Chinese hamster ovary cells.J Mol Biol. 1993 Aug 5;232(3):766-78. doi: 10.1006/jmbi.1993.1430. J Mol Biol. 1993. PMID: 8355269
-
HsOrc4-Dependent Dna Remodeling of the ori-β Dhfr Replicator.Cell Mol Biol Lett. 2015 Dec;20(4):549-61. doi: 10.1515/cmble-2015-0032. Cell Mol Biol Lett. 2015. PMID: 26124052
Cited by
-
Type D retrovirus Gag polyprotein interacts with the cytosolic chaperonin TRiC.J Virol. 2001 Mar;75(6):2526-34. doi: 10.1128/JVI.75.6.2526-2534.2001. J Virol. 2001. PMID: 11222675 Free PMC article.
-
Imaging of single hairpin ribozymes in solution by atomic force microscopy.RNA. 2001 Jun;7(6):887-95. doi: 10.1017/s1355838201002473. RNA. 2001. PMID: 11421363 Free PMC article.
-
The Chinese hamster dihydrofolate reductase replication origin beta is active at multiple ectopic chromosomal locations and requires specific DNA sequence elements for activity.Mol Cell Biol. 2001 Feb;21(4):1098-110. doi: 10.1128/MCB.21.4.1098-1110.2001. Mol Cell Biol. 2001. PMID: 11158297 Free PMC article.
-
Defined sequence modules and an architectural element cooperate to promote initiation at an ectopic mammalian chromosomal replication origin.Mol Cell Biol. 2004 May;24(10):4138-50. doi: 10.1128/MCB.24.10.4138-4150.2004. Mol Cell Biol. 2004. PMID: 15121836 Free PMC article.
-
Condensation by DNA looping facilitates transfer of large DNA molecules into mammalian cells.Nucleic Acids Res. 2001 May 1;29(9):1982-8. doi: 10.1093/nar/29.9.1982. Nucleic Acids Res. 2001. PMID: 11328883 Free PMC article.
References
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

