The specificity of protein-DNA crosslinking by formaldehyde: in vitro and in drosophila embryos
- PMID: 10606672
- PMCID: PMC102536
- DOI: 10.1093/nar/28.2.e4
The specificity of protein-DNA crosslinking by formaldehyde: in vitro and in drosophila embryos
Abstract
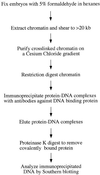
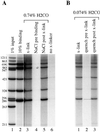
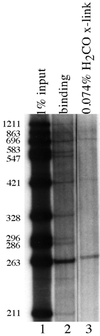
Formaldehyde crosslinking has been widely used to study binding of specific proteins to DNA elements in intact cells. However, previous studies have not determined if this crosslinker preserves the bona fide pattern of DNA binding. Here we show that formaldehyde crosslinking of Drosophila embryos maps an interaction of the transcription factor Zeste to a known target element in the Ultrabithorax promoter. This data agrees broadly with previous mapping of the same Zeste binding sites by in vivo UV crosslinking, though the formaldehyde method does give a low, possibly artifactual signal on other DNA fragments that is not detected by the UV method. We also demonstrate, using an in vitro assay, that formaldehyde crosslinking accurately reflects the DNA binding specificities of both Zeste and a second transcription factor, Eve. The crosslinking reagent methylene blue is shown to preserve DNA binding specificity in vitro as well. Our results suggest that crosslinking by formaldehyde, and possibly also by methylene blue, provide an accurate guide to the interaction of proteins with their high affinity target sites in cells.
Figures
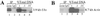


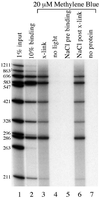
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

