Genetic diversity of clinical and environmental isolates of Vibrio cholerae determined by amplified fragment length polymorphism fingerprinting
- PMID: 10618216
- PMCID: PMC91798
- DOI: 10.1128/AEM.66.1.148-153.2000
Genetic diversity of clinical and environmental isolates of Vibrio cholerae determined by amplified fragment length polymorphism fingerprinting
Abstract
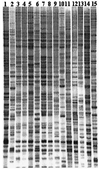
Vibrio cholerae, the causative agent of major epidemics of diarrheal disease in Bangladesh, South America, Southeastern Asia, and Africa, was isolated from clinical samples and from aquatic environments during and between epidemics over the past 20 years. To determine the evolutionary relationships and molecular diversity of these strains, in order to understand sources, origin, and epidemiology, a novel DNA fingerprinting technique, amplified fragment length polymorphism (AFLP), was employed. Two sets of restriction enzyme-primer combinations were tested for fingerprinting of V. cholerae serogroup O1, O139, and non-O1, O139 isolates. Amplification of HindIII- and TaqI-digested genomic DNA produced 30 to 50 bands for each strain. However, this combination, although capable of separating environmental isolates of O1 and non-O1 strains, was unable to distinguish between O1 and O139 clinical strains. This result confirmed that clinical O1 and O139 strains are genetically closely related. On the other hand, AFLP analyses of restriction enzyme ApaI- and TaqI-digested genomic DNA yielded 20 to 30 bands for each strain, but were able to separate O1 from O139 strains. Of the 74 strains examined with the latter combination, 26 serogroup O1 strains showed identical banding patterns and were represented by the O1 El Tor strain of the seventh pandemic. A second group, represented by O139 Bengal, included 12 strains of O139 clinical isolates, with 7 from Thailand, 3 from Bangladesh, and 2 from India. Interestingly, an O1 clinical isolate from Africa also grouped with the O139 clinical isolates. Eight clinical O1 isolates from Mexico grouped separately from the O1 El Tor of the seventh pandemic, suggesting an independent origin of these isolates. Identical fingerprints were observed between an O1 environmental isolate from a river in Chile and an O1 clinical strain from Kenya, both isolated more than 10 years apart. Both strains were distinct from the O1 seventh pandemic strain. Two O139 clinical isolates from Africa clustered with environmental non-O1 isolates, independent of other O139 strains included in the study. These results suggest that although a single clone of pathogenic V. cholerae appears responsible for many cases of cholera in Asia, Africa, and Latin America during the seventh pandemic, other cases of clinical cholera were caused by toxigenic V. cholerae strains that appear to have been derived locally from environmental O1 or non-O1 strains.
Figures


Similar articles
-
Molecular analysis of Vibrio cholerae O1, O139, non-O1, and non-O139 strains: clonal relationships between clinical and environmental isolates.Appl Environ Microbiol. 2001 Feb;67(2):910-21. doi: 10.1128/AEM.67.2.910-921.2001. Appl Environ Microbiol. 2001. PMID: 11157262 Free PMC article.
-
Emergence of a new clone of toxigenic Vibrio cholerae O1 biotype El Tor displacing V. cholerae O139 Bengal in Bangladesh.J Clin Microbiol. 1997 Mar;35(3):624-30. doi: 10.1128/jcm.35.3.624-630.1997. J Clin Microbiol. 1997. PMID: 9041401 Free PMC article.
-
Molecular analysis of toxigenic Vibrio cholerae O139 Bengal strains isolated in Bangladesh between 1993 and 1996: evidence for emergence of a new clone of the Bengal vibrios.J Clin Microbiol. 1997 Sep;35(9):2299-306. doi: 10.1128/jcm.35.9.2299-2306.1997. J Clin Microbiol. 1997. PMID: 9276406 Free PMC article.
-
Epidemiology & molecular biology of Vibrio cholerae O139 Bengal.Indian J Med Res. 1996 Jul;104:14-27. Indian J Med Res. 1996. PMID: 8783504 Review.
-
Non-serogroup O1/O139 agglutinable Vibrio cholerae: a phylogenetically and genealogically neglected yet emerging potential pathogen of clinical relevance.Arch Microbiol. 2022 May 14;204(6):323. doi: 10.1007/s00203-022-02866-1. Arch Microbiol. 2022. PMID: 35567650 Free PMC article. Review.
Cited by
-
Genomic diversity of clinical and environmental Vibrio cholerae strains isolated in Brazil between 1991 and 2001 as revealed by fluorescent amplified fragment length polymorphism analysis.J Clin Microbiol. 2003 May;41(5):1946-50. doi: 10.1128/JCM.41.5.1946-1950.2003. J Clin Microbiol. 2003. PMID: 12734232 Free PMC article.
-
Pandemic spread of cholera: genetic diversity and relationships within the seventh pandemic clone of Vibrio cholerae determined by amplified fragment length polymorphism.J Clin Microbiol. 2002 Jan;40(1):172-81. doi: 10.1128/JCM.40.1.172-181.2002. J Clin Microbiol. 2002. PMID: 11773113 Free PMC article.
-
Genetic diversity of Vibrio cholerae in Chesapeake Bay determined by amplified fragment length polymorphism fingerprinting.Appl Environ Microbiol. 2000 Jan;66(1):140-7. doi: 10.1128/AEM.66.1.140-147.2000. Appl Environ Microbiol. 2000. PMID: 10618215 Free PMC article.
-
Allelic diversity and population structure in Vibrio cholerae O139 Bengal based on nucleotide sequence analysis.J Bacteriol. 2002 Mar;184(5):1304-13. doi: 10.1128/JB.184.5.1304-1313.2002. J Bacteriol. 2002. PMID: 11844759 Free PMC article.
-
Phenotypic and genotypic characteristics and epidemiological significance of ctx+ strains of Vibrio cholerae isolated from seafood in Malaysia.Appl Environ Microbiol. 2004 Apr;70(4):1964-72. doi: 10.1128/AEM.70.4.1964-1972.2004. Appl Environ Microbiol. 2004. PMID: 15066786 Free PMC article.
References
-
- Arias C, Verdonck L, Swings J, Aznar R, Garay E. A polyphasic approach to study the intraspecific diversity amongst Vibrio vulnificus isolates. Syst Appl Microbiol. 1997;20:622–633.
-
- Ausubel F M, Brent R, Kingston R E, Moore D D, Seidman J G, Smith J A, Struhl K. Short protocols in molecular biology. 3rd ed. New York, N.Y: John Wiley & Sons, Inc.; 1995.
-
- Centers for Disease Control. Cholera—Peru, 1991. Morbid Mortal Weekly Rep. 1991;40:108–110. - PubMed
-
- Choudhury S R, Bhadra R K, Das J. Genome size and restriction fragment length polymorphism analysis of Vibrio cholerae strains belonging to different serovars and biotypes. FEMS Microbiol Lett. 1994;115:329–334. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

