Toxin-antitoxin modules may regulate synthesis of macromolecules during nutritional stress
- PMID: 10633087
- PMCID: PMC94316
- DOI: 10.1128/JB.182.3.561-572.2000
Toxin-antitoxin modules may regulate synthesis of macromolecules during nutritional stress
Figures
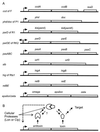




References
-
- Bahassi E M, O'Dea M H, Allali N, Messens J, Gellert M, Couturier M. Interactions of CcdB with DNA gyrase. Inactivation of GyrA, poisoning of the gyrase-DNA complex, and the antidote action of CcdA. J Biol Chem. 1999;274:10936–10944. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

