Expression profiling of the maize flavonoid pathway genes controlled by estradiol-inducible transcription factors CRC and P
- PMID: 10634908
- PMCID: PMC140215
- DOI: 10.1105/tpc.12.1.65
Expression profiling of the maize flavonoid pathway genes controlled by estradiol-inducible transcription factors CRC and P
Abstract
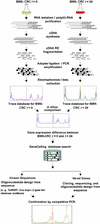
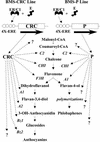
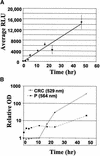
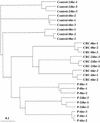
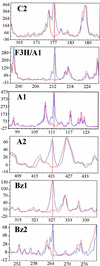
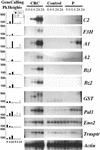
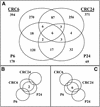
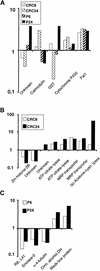
To determine the scope of gene expression controlled by the maize transcription factors C1/R and P, which are responsible for activating flavonoid synthesis, we used GeneCalling, an open-ended, gel-based, mRNA-profiling technology, to analyze cell suspension lines of the maize inbred Black Mexican Sweet (BMS) that harbored estradiol-inducible versions of these factors. BMS cells were transformed with a continually expressed estrogen receptor/maize C1 activator domain fusion gene (ER-C1) and either a fusion of C1 and R (CRC), P, or luciferase genes regulated by a promoter containing four repeats of an estrogen receptor binding site. Increasing amounts of luciferase activity, anthocyanins, and flavan-4-ols were detected in the respective cell lines after the addition of estradiol. The expression of both known and novel genes was detected simultaneously in these BMS lines by profiling the mRNA isolated from replicate samples at 0, 6, and 24 hr after estradiol treatment. Numerous cDNA fragments were identified that showed a twofold or greater difference in abundance at 6 and 24 hr than at 0 hr. The cDNA fragments from the known flavonoid genes, except chalcone isomerase (chi1), were induced in the CRC-expressing line after hormone induction, whereas only the chalcone synthase (c2) and flavanone/dihydroflavonol reductase (a1) genes were induced in the P-expressing line, as was expected. Many novel cDNA fragments were also induced or repressed by lines expressing CRC alone, P alone, or both transcription factors in unique temporal patterns. The temporal differences and the evidence of repression indicate a more diverse set of regulatory controls by CRC or P than originally expected. GeneCalling analysis was successful in detecting members of complex metabolic pathways and uncovering novel genes that were either coincidentally regulated or directly involved in such pathways.
Figures

References
-
- Ashton, A.R., Jenkins, C.L.D., and Whitfeld, P.R. (1994). Molecular cloning of two different cDNAs for maize acetyl-CoA carboxylase. Plant Mol. Biol. 24 35–49. - PubMed
-
- Avila, J., Nieto, C., Canas, L., Benito, M.J., and Paz-Ares, J. (1993). Petunia hybrida genes related to the maize regulatory C1 gene and to animal Myb proto-oncogenes. Plant J. 3 553–562. - PubMed
-
- Bachem, C.W., van der Hoeven, R.S., de Bruijn, S.M., Vreugdenhil, D., Zabeau, M., and Visser, R.G. (1996). Visualization of differential gene expression using a novel method of RNA fingerprinting based on AFLP: Analysis of gene expression during potato tuber development. Plant J. 9 745–753. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

