RNA double cleavage by a hairpin-derived twin ribozyme
- PMID: 10648779
- PMCID: PMC102578
- DOI: 10.1093/nar/28.4.886
RNA double cleavage by a hairpin-derived twin ribozyme
Abstract
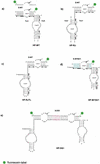
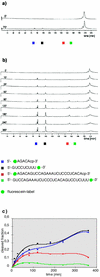
The hairpin ribozyme is a small catalytic RNA that catalyses reversible sequence-specific RNA hydrolysis in trans. It consists of two domains, which interact with each other by docking in an antiparallel fashion. There is a region between the two domains acting as a flexible hinge for interdomain interactions to occur. Hairpin ribozymes with reverse-joined domains have been constructed by dissecting the domains at the hinge and rejoining them in reverse order. We have used both the conventional and reverse-joined hairpin ribozymes for the design of a hairpin-derived twin ribozyme. We show that this twin ribozyme cleaves a suitable RNA substrate at two specific sites while maintaining the target specificity of the individual monoribozymes. For characterisation of the studied ribozymes we have evaluated a quantitative assay of sequence-specific ribozyme activity using fluorescently labelled RNA substrates in conjunction with an automated DNA sequencer. This assay was found to be applicable with hairpin and hairpin-derived ribozymes. The results demonstrate the potential of hairpin ribozymes for multi-target strategies of RNA cleavage and suggest the possibility for employing hairpin-derived twin ribozymes as powerful tools for RNA manipulation in vitro and in vivo.
Figures


Similar articles
-
Efficient RNA ligation by reverse-joined hairpin ribozymes and engineering of twin ribozymes consisting of conventional and reverse-joined hairpin ribozyme units.FEBS J. 2005 Sep;272(17):4464-74. doi: 10.1111/j.1742-4658.2005.04865.x. FEBS J. 2005. PMID: 16128815
-
Modification of primary structures of hairpin ribozymes for probing active conformations.J Mol Biol. 1995 Sep 22;252(3):296-304. doi: 10.1006/jmbi.1995.0497. J Mol Biol. 1995. PMID: 7563051
-
Kinetics and thermodynamics of intermolecular catalysis by hairpin ribozymes.Biochemistry. 1995 Dec 5;34(48):15813-28. doi: 10.1021/bi00048a027. Biochemistry. 1995. PMID: 7495813
-
The many faces of the hairpin ribozyme: structural and functional variants of a small catalytic RNA.IUBMB Life. 2012 Jan;64(1):36-47. doi: 10.1002/iub.575. Epub 2011 Nov 30. IUBMB Life. 2012. PMID: 22131309 Review.
-
The hairpin ribozyme: discovery, two-dimensional model, and development for gene therapy.Prog Nucleic Acid Res Mol Biol. 1998;58:1-39. doi: 10.1016/s0079-6603(08)60032-x. Prog Nucleic Acid Res Mol Biol. 1998. PMID: 9308362 Review.
Cited by
-
Synthesis of specifically modified oligonucleotides for application in structural and functional analysis of RNA.J Nucleic Acids. 2011;2011:805253. doi: 10.4061/2011/805253. Epub 2011 Oct 12. J Nucleic Acids. 2011. PMID: 22013508 Free PMC article.
-
Ribozyme suppression of endogenous thyroid hormone receptor activity in Xenopus laevis cells.Nucleic Acids Res. 2002 Aug 1;30(15):3490-6. doi: 10.1093/nar/gkf461. Nucleic Acids Res. 2002. PMID: 12140335 Free PMC article.
-
A continuous kinetic assay for RNA-cleaving deoxyribozymes, exploiting ethidium bromide as an extrinsic fluorescent probe.Nucleic Acids Res. 2002 Oct 15;30(20):e112. doi: 10.1093/nar/gnf111. Nucleic Acids Res. 2002. PMID: 12384614 Free PMC article.
-
Multivalent DNAzyme agents for cleaving folded RNA.Nucleic Acids Res. 2024 Jun 10;52(10):5866-5879. doi: 10.1093/nar/gkae295. Nucleic Acids Res. 2024. PMID: 38661191 Free PMC article.
-
Ligand-induced conformational capture of a synthetic tetracycline riboswitch revealed by pulse EPR.RNA. 2011 Jan;17(1):182-8. doi: 10.1261/rna.2222811. Epub 2010 Nov 19. RNA. 2011. PMID: 21097555 Free PMC article.
References
-
- Lo K.M.S., Biasolo,M.A., Dehni,G., Palu,G. and Haseltine,W.A. (1992) Virology, 190, 176–183. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

