Active residues and viral substrate cleavage sites of the protease of the birnavirus infectious pancreatic necrosis virus
- PMID: 10666235
- PMCID: PMC111686
- DOI: 10.1128/jvi.74.5.2057-2066.2000
Active residues and viral substrate cleavage sites of the protease of the birnavirus infectious pancreatic necrosis virus
Abstract
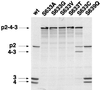
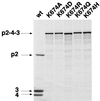
The polyprotein of infectious pancreatic necrosis virus (IPNV), a birnavirus, is processed by the viral protease VP4 (also named NS) to generate three polypeptides: pVP2, VP4, and VP3. Site-directed mutagenesis at 42 positions of the IPNV VP4 protein was performed to determine the active site and the important residues for the protease activity. Two residues (serine 633 and lysine 674) were critical for cleavage activity at both the pVP2-VP4 and the VP4-VP3 junctions. Wild-type activity at the pVP2-VP4 junction and a partial block (with an alteration of the cleavage specificity) at the VP4-VP3 junction were observed when replacement occurred at histidines 547 and 679. A similar observation was made when aspartic acid 693 was replaced by leucine, but wild-type activity and specificity were found when substituted by glutamine or asparagine. Sequence comparison between IPNV and two birnavirus (infectious bursal disease virus and Drosophila X virus) VP4s revealed that serine 633 and lysine 674 are conserved in these viruses, in contrast to histidines 547 and 679. The importance of serine 633 and lysine 674 is reminiscent of the protease active site of bacterial leader peptidases and their mitochondrial homologs and of the bacterial LexA-like proteases. Self-cleavage sites of IPNV VP4 were determined at the pVP2-VP4 and VP4-VP3 junctions by N-terminal sequencing and mutagenesis. Two alternative cleavage sites were also identified in the carboxyl domain of pVP2 by cumulative mutagenesis. The results suggest that VP4 cleaves the (Ser/Thr)-X-Ala / (Ser/Ala)-Gly motif, a target sequence with similarities to bacterial leader peptidases and herpesvirus protease cleavage sites.
Figures


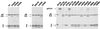





Similar articles
-
Role of Ser-652 and Lys-692 in the protease activity of infectious bursal disease virus VP4 and identification of its substrate cleavage sites.J Gen Virol. 2000 Apr;81(Pt 4):983-92. doi: 10.1099/0022-1317-81-4-983. J Gen Virol. 2000. PMID: 10725424
-
Characterization of cleavage sites and protease activity in the polyprotein precursor of Japanese marine aquabirnavirus and expression analysis of generated proteins by a VP4 protease activity in four distinct cell lines.Arch Virol. 2007;152(6):1103-14. doi: 10.1007/s00705-007-0935-z. Epub 2007 Mar 5. Arch Virol. 2007. PMID: 17334948
-
Crystal structure of the VP4 protease from infectious pancreatic necrosis virus reveals the acyl-enzyme complex for an intermolecular self-cleavage reaction.J Biol Chem. 2007 Aug 24;282(34):24928-37. doi: 10.1074/jbc.M701551200. Epub 2007 Jun 6. J Biol Chem. 2007. PMID: 17553791
-
Crystal structure of a novel viral protease with a serine/lysine catalytic dyad mechanism.J Mol Biol. 2006 May 19;358(5):1378-89. doi: 10.1016/j.jmb.2006.02.045. Epub 2006 Mar 6. J Mol Biol. 2006. PMID: 16584747
-
[Interaction mechanism of marine birnavirus (MABV) in fish cell lines].Uirusu. 2005 Jun;55(1):133-44. doi: 10.2222/jsv.55.133. Uirusu. 2005. PMID: 16308540 Review. Japanese.
Cited by
-
Blotched snakehead virus is a new aquatic birnavirus that is slightly more related to avibirnavirus than to aquabirnavirus.J Virol. 2003 Jan;77(1):719-25. doi: 10.1128/jvi.77.1.719-725.2003. J Virol. 2003. PMID: 12477876 Free PMC article.
-
Whole Genome Sequencing of Infectious Bursal Disease Viruses Isolated from a Californian Outbreak Unravels the Underlying Virulence Markers and Highlights Positive Selection Incidence.Viruses. 2023 Oct 3;15(10):2044. doi: 10.3390/v15102044. Viruses. 2023. PMID: 37896821 Free PMC article.
-
The Over-40-Years-Epidemic of Infectious Bursal Disease Virus in China.Viruses. 2022 Oct 14;14(10):2253. doi: 10.3390/v14102253. Viruses. 2022. PMID: 36298808 Free PMC article. Review.
-
Genome assembly and particle maturation of the birnavirus infectious pancreatic necrosis virus.J Virol. 2004 Dec;78(24):13829-38. doi: 10.1128/JVI.78.24.13829-13838.2004. J Virol. 2004. PMID: 15564491 Free PMC article.
-
Infectious pancreatic necrosis virus proteins VP2, VP3, VP4 and VP5 antagonize IFNa1 promoter activation while VP1 induces IFNa1.Virus Res. 2015 Jan 22;196:113-21. doi: 10.1016/j.virusres.2014.11.018. Epub 2014 Nov 24. Virus Res. 2015. PMID: 25445351 Free PMC article.
References
-
- Allaire M, Chernaia M M, Malcolm B A, James M N G. Picornaviral 3C cysteine proteinases have a fold similar to chymotrypsin-like serine proteinases. Nature. 1994;369:72–76. - PubMed
-
- Allsop A E, Ashby M, Brooks G, Bruton G, Coulton S, Edwards P D, Elsmere S A, Hatton I K, Kaura A C, McLean S D, Pearson M J, Pearson N D, Perry C R, Smale T, Southgate R. Inhibition of protein export in bacteria: the signalling of a new role for β-lactams. In: Bentley P H, O'Hanlon P J, editors. Anti-infectives: recent advances in chemistry and structure-activity relationships—1997. Cambridge, England: The Royal Society of Chemistry; 1997. pp. 61–72.
-
- Azad A A, Jagadish M N, Brown M A, Hudson P J. Deletion mapping and expression in Escherichia coli of the large genomic segment of a birnavirus. Virology. 1987;161:145–152. - PubMed
-
- Babé L A, Craik C S. Viral proteases: evolution of diverse structural motifs to optimize function. Cell. 1997;91:427–430. - PubMed
-
- Barrett A J, Rawlings N D. Families and clans of serine peptidases. Arch Biochem Biophys. 1995;318:247–250. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Miscellaneous

