Mutations in ribosomal protein L16 conferring reduced susceptibility to evernimicin (SCH27899): implications for mechanism of action
- PMID: 10681347
- PMCID: PMC89755
- DOI: 10.1128/AAC.44.3.732-738.2000
Mutations in ribosomal protein L16 conferring reduced susceptibility to evernimicin (SCH27899): implications for mechanism of action
Abstract
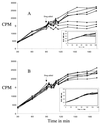
A clinical isolate of Streptococcus pneumoniae (SP#5) that showed decreased susceptibility to evernimicin (MIC, 1.5 microgram/ml) was investigated. A 4,255-bp EcoRI fragment cloned from SP#5 was identified by its ability to transform evernimicin-susceptible S. pneumoniae R6 (MIC, 0.03 microgram/ml) such that the evernimicin MIC was 1.5 microgram/ml. Nucleotide sequence analysis of this fragment revealed that it contained portions of the S10-spc ribosomal protein operons. The nucleotide sequences of resistant and susceptible isolates were compared, and a point mutation (thymine to guanine) that causes an Ile52-Ser substitution in ribosomal protein L16 was identified. The role of this mutation in decreasing susceptibility to evernimicin was confirmed by direct transformation of the altered L16 gene. The presence of the L16 mutation in the resistant strain suggests that evernimicin is an inhibitor of protein synthesis. This was confirmed by inhibition studies using radiolabeled substrates, which showed that the addition of evernimicin at sub-MIC levels resulted in a rapid decrease in the incorporation of radiolabeled isoleucine in a susceptible isolate (SP#3) but was much less effective against SP#5. The incorporation of isoleucine showed a linear response to the dose level of evernimicin. The incorporation of other classes of labeled substrates was unaffected or much delayed, indicating that these were secondary effects.
Figures


References
-
- Ausubel F M, Brent R, Kingston R E, Moore D D, Seidman J G, Smith J A, Struhl K, editors. Current protocols in molecular biology. New York, N.Y: John Wiley and Sons, Inc.; 1995.
-
- Bernabeu C, Vázquez D, Ballesta J P G. The involvement of protein L16 on ribosomal peptidyl transferase activity. Eur J Biochem. 1977;79:469–472. - PubMed
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

