Yeast heterochromatin is a dynamic structure that requires silencers continuously
- PMID: 10691737
- PMCID: PMC316382
Yeast heterochromatin is a dynamic structure that requires silencers continuously
Abstract
Transcriptional silencing of the HM loci in yeast requires cis-acting elements, termed silencers, that function during S-phase passage to establish the silent state. To study the role of the regulatory elements in maintenance of repression, site-specific recombination was used to uncouple preassembled silent chromatin fragments from silencers. DNA rings excised from HMR were initially silent but ultimately reactivated, even in G(1)- or G(2)/M-arrested cells. In contrast, DNA rings bearing HML-derived sequence were stably repressed due to the presence of a protosilencing element. These data show that silencers (or protosilencers) are required continuously for maintenance of silent chromatin. Reactivation of unstably repressed rings was blocked by overexpression of silencing proteins Sir3p and Sir4p, and chromatin immunoprecipitation studies showed that overexpressed Sir3p was incorporated into silent chromatin. Importantly, the protein was incorporated even when expressed outside of S phase, during G(1) arrest. That silencing factors can associate with and stabilize preassembled silent chromatin in non-S-phase cells demonstrates that heterochromatin in yeast is dynamic.
Figures
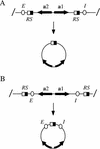
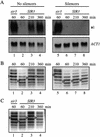





References
-
- Andrulis E, Neiman AM, Zappulla DC, Sternglanz R. Perinuclear localization of chromatin facilitates transcriptional silencing. Nature. 1998;394:592–595. - PubMed
-
- Aparicio OM, Gottschling DE. Overcoming telomeric silencing: A trans-activator competes to establish gene expression in a cell-cycle dependent way. Genes & Dev. 1994;8:1133–1146. - PubMed
-
- Aparicio OM, Billington BL, Gottschling DE. Modifiers of position effect are shared between telomeric and silent mating-type loci in Saccharomyces cerevisiae. Cell. 1991;66:1279–1287. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
