Semaphorin SEMA3F localization in malignant human lung and cell lines: A suggested role in cell adhesion and cell migration
- PMID: 10702410
- PMCID: PMC1876858
- DOI: 10.1016/S0002-9440(10)64962-0
Semaphorin SEMA3F localization in malignant human lung and cell lines: A suggested role in cell adhesion and cell migration
Abstract
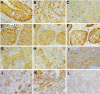
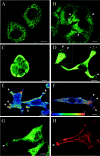
Semaphorins/collapsins are a family of secreted and membrane-associated proteins involved in nerve growth cone migration. However, some are expressed widely in adult tissues suggesting additional functions. SEMA3F/H.SemaIV was previously isolated from a 3p21.3 homozygous deletion region in human lung cancer. We studied SEMA3F cellular localization using our previously characterized anti-SEMA3F antibody. In normal lung, SEMA3F was found in all epithelial cells at the cytoplasmic membrane and, to a lesser extent, in the cytoplasm. In lung tumors, the localization was predominantly cytoplasmic, and the levels were comparatively reduced. In non-small-cell lung carcinomas, low levels correlated with higher stage. In all tumors, an exclusive cytoplasmic localization of SEMA3F correlated with high levels of vascular endothelial growth factor and was related to the grade and aggressiveness. This suggests that vascular endothelial growth factor might compete with SEMA3F for binding to their common receptors, neuropilin-1 and -2 and might contribute to SEMA3F delocalization and deregulation in lung tumor. In parallel studies, SEMA3F distribution was examined in cell cultures by confocal microscopy. Marked staining was observed in pseudopods and in the leading edge or ruffling membranes of lamellipods or cellular protrusions in motile cells. SEMA3F was also observed at the interface of adjacent interacting cells suggesting a role in cell motility and cell adhesion.
Figures

Similar articles
-
Expression of VEGF, semaphorin SEMA3F, and their common receptors neuropilins NP1 and NP2 in preinvasive bronchial lesions, lung tumours, and cell lines.J Pathol. 2003 Jul;200(3):336-47. doi: 10.1002/path.1367. J Pathol. 2003. PMID: 12845630
-
Semaphorin SEMA3F and VEGF have opposing effects on cell attachment and spreading.Neoplasia. 2003 Jan-Feb;5(1):83-92. doi: 10.1016/s1476-5586(03)80020-9. Neoplasia. 2003. PMID: 12659673 Free PMC article.
-
Selective suppression of in vivo tumorigenicity by semaphorin SEMA3F in lung cancer cells.Neoplasia. 2005 May;7(5):457-65. doi: 10.1593/neo.04721. Neoplasia. 2005. PMID: 15967098 Free PMC article.
-
Neuropilin and its ligands in normal lung and cancer.Adv Exp Med Biol. 2002;515:103-14. doi: 10.1007/978-1-4615-0119-0_9. Adv Exp Med Biol. 2002. PMID: 12613547 Review.
-
Ectopic hormones in lung cancer.Ergeb Inn Med Kinderheilkd. 1984;53:133-64. doi: 10.1007/978-3-642-69841-5_5. Ergeb Inn Med Kinderheilkd. 1984. PMID: 6094177 Review. No abstract available.
Cited by
-
Semaphorin 5A suppresses the proliferation and migration of lung adenocarcinoma cells.Int J Oncol. 2020 Jan;56(1):165-177. doi: 10.3892/ijo.2019.4932. Epub 2019 Dec 2. Int J Oncol. 2020. PMID: 31789397 Free PMC article.
-
Response to Hypoxia and the Ensuing Dysregulation of Inflammation Impacts Mycobacterium tuberculosis Pathogenicity.Am J Respir Crit Care Med. 2022 Jul 1;206(1):94-104. doi: 10.1164/rccm.202112-2747OC. Am J Respir Crit Care Med. 2022. PMID: 35412961 Free PMC article.
-
Expression of semaphorin 3A and neuropilin 1 in asthma.J Korean Med Sci. 2013 Oct;28(10):1435-42. doi: 10.3346/jkms.2013.28.10.1435. Epub 2013 Sep 25. J Korean Med Sci. 2013. PMID: 24133346 Free PMC article.
-
Inhibition of lung cancer cell growth and induction of apoptosis after reexpression of 3p21.3 candidate tumor suppressor gene SEMA3B.Proc Natl Acad Sci U S A. 2001 Nov 20;98(24):13954-9. doi: 10.1073/pnas.231490898. Proc Natl Acad Sci U S A. 2001. PMID: 11717452 Free PMC article.
-
Secreted semaphorin 5A suppressed pancreatic tumour burden but increased metastasis and endothelial cell proliferation.Br J Cancer. 2012 Jul 24;107(3):501-7. doi: 10.1038/bjc.2012.298. Epub 2012 Jul 10. Br J Cancer. 2012. PMID: 22782341 Free PMC article.
References
-
- : Semaphorin Nomenclature Committee: Unified nomenclature for the semaphorins/collapsins. Cell 1999, 97:551-552 - PubMed
-
- Roche J, Boldog F, Robinson M, Robinson L, Varella-Garcia M, Swanton M, Waggoner B, Fishel R, Franklin W, Gemmill R, Drabkin H: Distinct 3p21.3 deletions in lung cancer, analysis of deleted genes, and identification of a new human semaphorin. Oncogene 1996, 12:1289-1297 - PubMed
-
- Xiang R, Hensel C, Garcia D, Carlson H, Kok K, Daly M, Kerbacher K, Van Den Berg A, Veldhuis P, Buys C, Naylor S: Isolation of the human semaphorin III/F gene (SEMA3F) at chromosome 3p21, a region deleted in lung cancer. Genomics 1996, 32:39-48 - PubMed
-
- Luo Y, Raible D, Raper A: Collapsin: a protein in brain that induces the collapse and paralysis of neuronal growth cones. Cell 1993, 75:217-227 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

