Asymmetric subunit organization of heterodimeric Rous sarcoma virus reverse transcriptase alphabeta: localization of the polymerase and RNase H active sites in the alpha subunit
- PMID: 10708441
- PMCID: PMC111825
- DOI: 10.1128/jvi.74.7.3245-3252.2000
Asymmetric subunit organization of heterodimeric Rous sarcoma virus reverse transcriptase alphabeta: localization of the polymerase and RNase H active sites in the alpha subunit
Abstract
The genes encoding the alpha (63-kDa) and beta (95-kDa) subunits of Rous sarcoma virus (RSV) reverse transcriptase (RT) or the entire Pol polypeptide (99 kDa) were mutated in the conserved aspartic acid residue Asp 181 of the polymerase active site (YMDD) or in the conserved Asp 505 residue of the RNase H active site. We have analyzed heterodimeric recombinant RSV alphabeta and alphaPol RTs within which one subunit was selectively mutated. When alphabeta heterodimers contained the Asp 181-->Asn mutation in their beta subunits, about 42% of the wild-type polymerase activity was detected, whereas when the heterodimers contained the same mutation in their alpha subunits, only 7.5% of the wild-type polymerase activity was detected. Similar results were obtained when the conserved Asp 505 residue of the RNase H active site was mutated to Asn. RNase H activity was clearly detectable in alphabeta heterodimers mutated in the beta subunit but was lost when the mutation was present in the alpha subunit. In summary, our data imply that the polymerase and RNase H active sites are located in the alpha subunit of the heterodimeric RSV RT alphabeta.
Figures


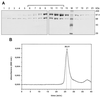






Similar articles
-
Requirements for minus-strand transfer catalyzed by Rous sarcoma virus reverse transcriptase.J Virol. 2001 Nov;75(21):10132-8. doi: 10.1128/JVI.75.21.10132-10138.2001. J Virol. 2001. PMID: 11581381 Free PMC article.
-
Homodimeric reverse transcriptases from rous sarcoma virus mutated within the polymerase or RNase H active site of one subunit are active.Eur J Biochem. 2000 Aug;267(15):4740-4. doi: 10.1046/j.1432-1327.2000.01530.x. Eur J Biochem. 2000. PMID: 10903507
-
Soluble Rous sarcoma virus reverse transcriptases alpha, alphabeta, and beta purified from insect cells are processive DNA polymerases that lack an RNase H 3' --> 5' directed processing activity.J Biol Chem. 1999 Sep 10;274(37):26329-36. doi: 10.1074/jbc.274.37.26329. J Biol Chem. 1999. PMID: 10473589
-
Isolation and characterization of Rous sarcoma virus recombinant reverse transcriptase dimers.Biochemistry (Mosc). 1999 Aug;64(8):933-7. Biochemistry (Mosc). 1999. PMID: 10498811
-
Subcellular localization and integration activities of rous sarcoma virus reverse transcriptase.J Virol. 2002 Jun;76(12):6205-12. doi: 10.1128/jvi.76.12.6205-6212.2002. J Virol. 2002. PMID: 12021354 Free PMC article.
Cited by
-
Role of integrase in reverse transcription of the Saccharomyces cerevisiae retrotransposon Ty1.Eukaryot Cell. 2005 Jun;4(6):1057-65. doi: 10.1128/EC.4.6.1057-1065.2005. Eukaryot Cell. 2005. PMID: 15947198 Free PMC article.
-
The role of template-primer in protection of reverse transcriptase from thermal inactivation.Nucleic Acids Res. 2002 Jul 15;30(14):3118-29. doi: 10.1093/nar/gkf417. Nucleic Acids Res. 2002. PMID: 12136094 Free PMC article.
-
Retroviral DNA Transposition: Themes and Variations.Microbiol Spectr. 2014 Dec;2(5):MDNA300052014. doi: 10.1128/microbiolspec.MDNA3-0005-2014. Microbiol Spectr. 2014. PMID: 25844274 Free PMC article.
-
Requirements for minus-strand transfer catalyzed by Rous sarcoma virus reverse transcriptase.J Virol. 2001 Nov;75(21):10132-8. doi: 10.1128/JVI.75.21.10132-10138.2001. J Virol. 2001. PMID: 11581381 Free PMC article.
-
Alternate polypurine tracts (PPTs) affect the rous sarcoma virus RNase H cleavage specificity and reveal a preferential cleavage following a GA dinucleotide sequence at the PPT-U3 junction.J Virol. 2005 Nov;79(21):13694-704. doi: 10.1128/JVI.79.21.13694-13704.2005. J Virol. 2005. PMID: 16227289 Free PMC article.
References
-
- Delarue M, Poch O, Tordo N, Moras D, Argos P. An attempt to unify the structure of polymerases. Protein Eng. 1990;3:461–467. - PubMed
-
- DeStefano J J, Wu W, Seehra J, McCoy J, Laston D, Albone E, Fay P J, Bambara R A. Characterization of an RNase H deficient mutant of human immunodeficiency virus-1 reverse transcriptase having an aspartate to asparagine change at position 498. Biochim Biophys Acta. 1994;1219:380–388. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

