BRCA1 interaction with RNA polymerase II reveals a role for hRPB2 and hRPB10alpha in activated transcription
- PMID: 10725406
- PMCID: PMC16207
- DOI: 10.1073/pnas.97.7.3148
BRCA1 interaction with RNA polymerase II reveals a role for hRPB2 and hRPB10alpha in activated transcription
Abstract
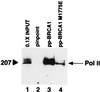
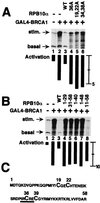
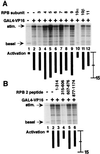
The functions of most of the 12 subunits of the RNA polymerase II (Pol II) enzyme are unknown. In this study, we demonstrate that two of the subunits, hRPB2 and hRPB10alpha, mediate the regulated stimulation of transcription. We find that the transcriptional coactivator BRCA1 interacts directly with the core Pol II complex in vitro. We tested whether single subunits from Pol II would compete with the intact Pol II complex to inhibit transcription stimulated by BRCA1. Excess purified Pol II subunits hRPB2 or hRPB10alpha blocked BRCA1- and VP16-dependent transcriptional activation in vitro with minimal effect on basal transcription. No other Pol II subunits tested inhibited activated transcription in these assays. Furthermore, hRPB10alpha, but not hRPB2, blocked Sp1-dependent activation.
Figures



Similar articles
-
BRCA1 associates with processive RNA polymerase II.J Biol Chem. 2003 Dec 26;278(52):52012-20. doi: 10.1074/jbc.M308418200. Epub 2003 Sep 23. J Biol Chem. 2003. PMID: 14506230
-
Semen Exosomes Promote Transcriptional Silencing of HIV-1 by Disrupting NF-κB/Sp1/Tat Circuitry.J Virol. 2018 Oct 12;92(21):e00731-18. doi: 10.1128/JVI.00731-18. Print 2018 Nov 1. J Virol. 2018. PMID: 30111566 Free PMC article.
-
BRCA1 cooperates with NUFIP and P-TEFb to activate transcription by RNA polymerase II.Oncogene. 2004 Jul 8;23(31):5316-29. doi: 10.1038/sj.onc.1207684. Oncogene. 2004. PMID: 15107825
-
Cofactor requirements for transcriptional activation by Sp1.Cold Spring Harb Symp Quant Biol. 1998;63:189-99. doi: 10.1101/sqb.1998.63.189. Cold Spring Harb Symp Quant Biol. 1998. PMID: 10384283 Review. No abstract available.
-
BRCA1: exploring the links to transcription.Trends Biochem Sci. 2000 Oct;25(10):469-74. doi: 10.1016/s0968-0004(00)01632-7. Trends Biochem Sci. 2000. PMID: 11050427 Review.
Cited by
-
Subcellular localization of RPB5-mediating protein and its putative functional partner.Mol Cell Biol. 2004 Oct;24(19):8556-66. doi: 10.1128/MCB.24.19.8556-8566.2004. Mol Cell Biol. 2004. PMID: 15367675 Free PMC article.
-
BRCA1 protein and nucleolin colocalize in breast carcinoma tissue and cancer cell lines.Am J Pathol. 2010 Mar;176(3):1203-14. doi: 10.2353/ajpath.2010.081063. Epub 2010 Jan 14. Am J Pathol. 2010. PMID: 20075200 Free PMC article.
-
Characterization of Arabidopsis thaliana ortholog of the human breast cancer susceptibility gene 1: AtBRCA1, strongly induced by gamma rays.Nucleic Acids Res. 2003 Feb 15;31(4):1148-55. doi: 10.1093/nar/gkg202. Nucleic Acids Res. 2003. PMID: 12582233 Free PMC article.
-
BRCA1 and Oxidative Stress.Cancers (Basel). 2014 Apr 3;6(2):771-95. doi: 10.3390/cancers6020771. Cancers (Basel). 2014. PMID: 24704793 Free PMC article.
-
Rpb3 promotes hepatocellular carcinoma through its N-terminus.Oncotarget. 2014 Oct 15;5(19):9256-68. doi: 10.18632/oncotarget.2389. Oncotarget. 2014. PMID: 25211001 Free PMC article.
References
-
- Buratowski S, Hahn S, Guarente L, Sharp P A. Cell. 1989;56:549–561. - PubMed
-
- Orphanides G, Lagrange T, Reinberg D. Genes Dev. 1996;10:2657–2683. - PubMed
-
- Parvin J D, Young R A. Curr Opin Genet Dev. 1998;8:565–570. - PubMed
-
- Meisterernst M, Roy A L, Lieu H M, Roeder R G. Cell. 1991;66:981–993. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

