Dysregulation of cyclin E gene expression in human cytomegalovirus-infected cells requires viral early gene expression and is associated with changes in the Rb-related protein p130
- PMID: 10756032
- PMCID: PMC111934
- DOI: 10.1128/jvi.74.9.4192-4206.2000
Dysregulation of cyclin E gene expression in human cytomegalovirus-infected cells requires viral early gene expression and is associated with changes in the Rb-related protein p130
Abstract
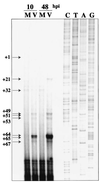
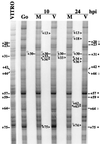
We have previously shown that many cell cycle regulatory gene products are markedly affected by infection of primary fibroblasts with human cytomegalovirus (HCMV) (F. M. Jault, J. M. Jault, F. Ruchti, E. A. Fortunato, C. Clark, J. Corbeil, D. D. Richman, and D. H. Spector, J. Virol. 69:6697-6704, 1995). One of these proteins, cyclin E, is a key determinant of cell cycle progression during G(1), and its mRNA levels are significantly increased in HCMV-infected fibroblasts (B. S. Salvant, E. A. Fortunato, and D. H. Spector, J. Virol. 72:3729-3741, 1998). To determine the molecular basis of this effect, we have examined the events that occur at the endogenous cyclin E promoter during the course of infection. In vivo dimethyl sulfate footprinting of the cyclin E promoter revealed several regions of protection and hypersensitivity that were unique to infected cells. In accord with this observation, we find that the virus-induced cyclin E transcripts initiate downstream of the start site identified in mock-infected cells, in regions where these newly appearing protected and hypersensitive sites occur. Viral gene expression is required for this induction. However, the viral immediate-early proteins IE1-72 and IE2-86, either alone or in combination, cannot induce expression of the endogenous cyclin E. The virus must progress past the immediate-early phase and express an early gene product(s) for activation of cyclin E expression. Moreover, IE1-72 does not appear to be required, as infection of cells with an HCMV mutant containing a deletion in the IE1-72 gene leads to full upregulation of cyclin E expression. Using electrophoretic mobility shift assays with infected cell extracts and a region of the cyclin E promoter that includes two previously defined E2F sites as the probe, we detected the appearance of an infection-specific banding pattern. One of the infection-specific bands contained the proteins E2F-4, DP-1, and p130, which were maintained in the infected cells as uniquely phosphorylated species. These results suggest that an altered E2F-4-DP-1-p130 complex along with viral early gene expression may play a role in the transcriptional regulation of cyclin E mRNA during HCMV infection.
Figures










Similar articles
-
Cell cycle dysregulation by human cytomegalovirus: influence of the cell cycle phase at the time of infection and effects on cyclin transcription.J Virol. 1998 May;72(5):3729-41. doi: 10.1128/JVI.72.5.3729-3741.1998. J Virol. 1998. PMID: 9557655 Free PMC article.
-
Disruption of PML-associated nuclear bodies by IE1 correlates with efficient early stages of viral gene expression and DNA replication in human cytomegalovirus infection.Virology. 2000 Aug 15;274(1):39-55. doi: 10.1006/viro.2000.0448. Virology. 2000. PMID: 10936087
-
The cyclin E promoter is activated by human cytomegalovirus 86-kDa immediate early protein.J Biol Chem. 1998 Aug 21;273(34):22075-82. doi: 10.1074/jbc.273.34.22075. J Biol Chem. 1998. PMID: 9705351
-
The 72K IE1 and 80K IE2 proteins of human cytomegalovirus independently trans-activate the c-fos, c-myc and hsp70 promoters via basal promoter elements.J Gen Virol. 1992 Sep;73 ( Pt 9):2385-93. doi: 10.1099/0022-1317-73-9-2385. J Gen Virol. 1992. PMID: 1328493
-
Early viral gene expression and function.In: Arvin A, Campadelli-Fiume G, Mocarski E, Moore PS, Roizman B, Whitley R, Yamanishi K, editors. Human Herpesviruses: Biology, Therapy, and Immunoprophylaxis. Cambridge: Cambridge University Press; 2007. Chapter 18. In: Arvin A, Campadelli-Fiume G, Mocarski E, Moore PS, Roizman B, Whitley R, Yamanishi K, editors. Human Herpesviruses: Biology, Therapy, and Immunoprophylaxis. Cambridge: Cambridge University Press; 2007. Chapter 18. PMID: 21348120 Free Books & Documents. Review.
Cited by
-
Tumor control in a model of bone marrow transplantation and acute liver-infiltrating B-cell lymphoma: an unpredicted novel function of cytomegalovirus.J Virol. 2002 Mar;76(6):2857-70. doi: 10.1128/jvi.76.6.2857-2870.2002. J Virol. 2002. PMID: 11861853 Free PMC article.
-
Mutation of glutamine to arginine at position 548 of IE2 86 in human cytomegalovirus leads to decreased expression of IE2 40, IE2 60, UL83, and UL84 and increased transcription of US8-9 and US29-32.J Virol. 2011 Nov;85(21):11098-110. doi: 10.1128/JVI.05315-11. Epub 2011 Aug 24. J Virol. 2011. PMID: 21865379 Free PMC article.
-
Biologic and immunologic effects of knockout of human cytomegalovirus pp65 nuclear localization signal.Clin Vaccine Immunol. 2009 Jun;16(6):935-43. doi: 10.1128/CVI.00011-09. Epub 2009 Apr 15. Clin Vaccine Immunol. 2009. PMID: 19369477 Free PMC article.
-
Human cytomegalovirus-encoded viral cyclin-dependent kinase (v-CDK) UL97 phosphorylates and inactivates the retinoblastoma protein-related p107 and p130 proteins.J Biol Chem. 2017 Apr 21;292(16):6583-6599. doi: 10.1074/jbc.M116.773150. Epub 2017 Mar 13. J Biol Chem. 2017. PMID: 28289097 Free PMC article.
-
Human cytomegalovirus RNA2.7 inhibits RNA polymerase II (Pol II) Serine-2 phosphorylation by reducing the interaction between Pol II and phosphorylated cyclin-dependent kinase 9 (pCDK9).Virol Sin. 2022 Jun;37(3):358-369. doi: 10.1016/j.virs.2022.02.011. Epub 2022 Feb 28. Virol Sin. 2022. PMID: 35537980 Free PMC article.
References
-
- Ausubel F M, et al. Current protocols in molecular biology. New York, N.Y: Greene Publishing Associates and Wiley-Interscience; 1987.
-
- Beijersbergen R L, Bernards R. Cell cycle regulation by the retinoblastoma family of growth inhibitory proteins. Biochim Biophys Acta. 1996;1287:103–120. - PubMed
-
- Bresnahan W A, Albrecht T, Thompson E A. The cyclin E promoter is activated by human cytomegalovirus 86-kDa immediate early protein. J Biol Chem. 1998;273:22075–22082. - PubMed
-
- Bresnahan W A, Boldogh I, Chi P, Thompson E A, Albrecht T. Inhibition of cellular Cdk2 activity blocks human cytomegalovirus replication. Virology. 1997;231:239–247. - PubMed
-
- Bresnahan W A, Boldogh I, Thompson E A, Albrecht T. Human cytomegalovirus inhibits cellular DNA synthesis and arrests productively infected cells in late G1. Virology. 1996;224:150–160. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

