AFM characterization of single strand-specific endonuclease activity on linear DNA
- PMID: 10756206
- PMCID: PMC103312
- DOI: 10.1093/nar/28.9.e39
AFM characterization of single strand-specific endonuclease activity on linear DNA
Abstract
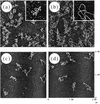
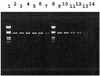
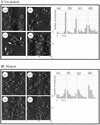
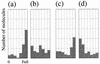
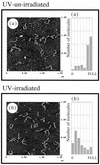
The specificity of nucleases for nicked and un-nicked double-stranded DNA has been characterized using atomic force microscopy (AFM). We have found that AFM has advantages over the usual macroscopic analyses, such as sucrose gradient centrifugation or electrophoresis, in characterizing nuclease digestion. In particular, short DNA fragments resulting from non-specific digestion were detected and, thus, the true length distribution of digested DNA was revealed. A simple numerical method is proposed to estimate the number of nicked sites per DNA molecule based on AFM images.
Figures
Similar articles
-
Induction of double-strand breaks by S1 nuclease, mung bean nuclease and nuclease P1 in DNA containing abasic sites and nicks.Nucleic Acids Res. 1995 Oct 11;23(19):3805-9. doi: 10.1093/nar/23.19.3805. Nucleic Acids Res. 1995. PMID: 7479020 Free PMC article.
-
[The action of S1 nuclease and a cloning strategy for microcircular DNAs].Sheng Wu Gong Cheng Xue Bao. 2003 Mar;19(2):240-3. Sheng Wu Gong Cheng Xue Bao. 2003. PMID: 15966330 Chinese.
-
A sizing artifact of DNA restriction fragments with unusually long single-stranded termini.Gene. 1991 Jun 15;102(1):79-81. doi: 10.1016/0378-1119(91)90542-j. Gene. 1991. PMID: 1650735
-
Analysis of radiation damage of DNA by atomic force microscopy in comparison with agarose gel electrophoresis studies.J Biochem Biophys Methods. 2000 Jul 10;44(1-2):31-40. doi: 10.1016/s0165-022x(00)00049-x. J Biochem Biophys Methods. 2000. PMID: 10889274
-
Assays for restriction endonucleases using plasmid substrates.Methods Mol Biol. 1994;30:385-96. doi: 10.1385/0-89603-256-6:385. Methods Mol Biol. 1994. PMID: 8004211 No abstract available.
Cited by
-
Molecular characterization of a novel mycovirus in the cultivated mushroom, Lentinula edodes.Virol J. 2012 Mar 6;9:60. doi: 10.1186/1743-422X-9-60. Virol J. 2012. PMID: 22390839 Free PMC article.
-
Direct real-time molecular scale visualisation of the degradation of condensed DNA complexes exposed to DNase I.Nucleic Acids Res. 2003 Jul 15;31(14):4001-5. doi: 10.1093/nar/gkg462. Nucleic Acids Res. 2003. PMID: 12853616 Free PMC article.
-
Modeling and Analysis of Intercalant Effects on Circular DNA Conformation.ACS Nano. 2016 Sep 27;10(9):8910-7. doi: 10.1021/acsnano.6b04876. Epub 2016 Aug 29. ACS Nano. 2016. PMID: 27559753 Free PMC article.
References
-
- Arscott P.G. and Bloomfield,V.A. (1992) Methods Enzymol., 211, 491–502. - PubMed
-
- Bustamante C., Keller,D. and Yang,G. (1993) Curr. Opin. Struct. Biol., 3, 363–372.
-
- Bustamante C., Erie,D.A. and Keller,D. (1994) Curr. Opin. Struct. Biol., 4, 750–760.
-
- Engel A., (1991) Annu. Rev. Biophys. Chem., 20, 79–108. - PubMed
-
- Hansma H.G. and Hoh,J. (1994) Annu. Rev. Biophys. Biochem. Struct., 23, 115–139. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

