Adeno-associated virus site-specifically integrates into a muscle-specific DNA region
- PMID: 10758163
- PMCID: PMC18323
- DOI: 10.1073/pnas.080079397
Adeno-associated virus site-specifically integrates into a muscle-specific DNA region
Abstract
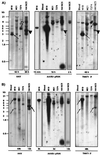
The nonpathogenic human virus adeno-associated virus type 2 (AAV) has evolved the potentially unique strategy to establish latency by site-specifically integrating its genome into human chromosome 19 (19q13.3-qter) at a locus designated AAVS1. This nonhomologous, site-specific recombination of viral DNA with the human genome provides a basis for developing targeted gene therapy vectors. To assess whether the region surrounding AAVS1 might have contributed to the selection of the specific integration site, we have investigated this locus. Here, we show that AAVS1 is closely linked to the slow skeletal troponin T gene, TNNT1, which has been mapped previously to 19q13.4. In support of this idea, we demonstrate that site-specific AAV DNA integration can result in the formation of TNNT1-AAV junctions. The question now arises whether muscle represents a natural target tissue for latent AAV infection. This possibility is of additional interest in view of recent observations that muscle tissue is particularly well suited for AAV-mediated gene transfer. The question also occurs whether latent infection by AAV can lead to phenotypic changes of the multinucleated muscle fiber cells.
Figures


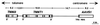
Similar articles
-
Adeno-associated virus site-specific integration and AAVS1 disruption.J Virol. 2004 Aug;78(15):7874-82. doi: 10.1128/JVI.78.15.7874-7882.2004. J Virol. 2004. PMID: 15254160 Free PMC article.
-
Site-specific integration of adeno-associated virus-based plasmid vectors in lipofected HeLa cells.Virology. 2000 Mar 15;268(2):391-401. doi: 10.1006/viro.1999.0122. Virology. 2000. PMID: 10704347
-
Recombinant junctions formed by site-specific integration of adeno-associated virus into an episome.J Virol. 1995 Nov;69(11):6917-24. doi: 10.1128/JVI.69.11.6917-6924.1995. J Virol. 1995. PMID: 7474109 Free PMC article.
-
Site-specific integration by adeno-associated virus.Proc Natl Acad Sci U S A. 1996 Oct 15;93(21):11288-94. doi: 10.1073/pnas.93.21.11288. Proc Natl Acad Sci U S A. 1996. PMID: 8876128 Free PMC article. Review.
-
Integration of adeno-associated virus (AAV) and recombinant AAV vectors.Annu Rev Genet. 2004;38:819-45. doi: 10.1146/annurev.genet.37.110801.143717. Annu Rev Genet. 2004. PMID: 15568995 Review.
Cited by
-
Relative influence of the adeno-associated virus (AAV) type 2 p5 element for recombinant AAV vector site-specific integration.J Virol. 2008 Mar;82(5):2590-3. doi: 10.1128/JVI.01956-07. Epub 2007 Dec 19. J Virol. 2008. PMID: 18094174 Free PMC article.
-
Rescue of the adeno-associated virus genome from a plasmid vector: evidence for rescue by replication.J Virol. 2003 Nov;77(21):11480-90. doi: 10.1128/jvi.77.21.11480-11490.2003. J Virol. 2003. PMID: 14557633 Free PMC article.
-
Substitution of adeno-associated virus Rep protein binding and nicking sites with human chromosome 19 sequences.Virol J. 2010 Sep 8;7:218. doi: 10.1186/1743-422X-7-218. Virol J. 2010. PMID: 20825662 Free PMC article.
-
Recombination in eukaryotic single stranded DNA viruses.Viruses. 2011 Sep;3(9):1699-738. doi: 10.3390/v3091699. Epub 2011 Sep 13. Viruses. 2011. PMID: 21994803 Free PMC article. Review.
-
Adeno-associated virus: a key to the human genome?Future Virol. 2010 Sep 1;5(5):555-574. doi: 10.2217/fvl.10.48. Future Virol. 2010. PMID: 21212830 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

