High spontaneous mutation rate in the hyperthermophilic archaeon Sulfolobus solfataricus is mediated by transposable elements
- PMID: 10762261
- PMCID: PMC111323
- DOI: 10.1128/JB.182.9.2574-2581.2000
High spontaneous mutation rate in the hyperthermophilic archaeon Sulfolobus solfataricus is mediated by transposable elements
Abstract
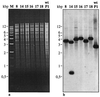
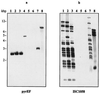
We have isolated uracil-auxotrophic mutants of the hyperthermophilic archaeon Sulfolobus solfataricus in order to explore the genomic stability and mutational frequencies of this organism and to identify complementable recipients for a selectable genetic transformation system. Positive selection of spontaneous mutants resistant to 5-fluoroorotate yielded uracil auxotrophs with frequencies of between 10(-4) and 10(-5) per sensitive, viable cell. Four different, nonhomologous insertion sequences (ISs) were identified at different positions within the chromosomal pyrEF locus of these mutants. They ranged in size from 1,058 to 1,439 bp and possessed properties typical of known transposable elements, i.e., terminal inverted repeats, flanking duplicated target sequences, and putative transposase genes encoding motifs that are indicative of the IS4-IS5 IS element families. Between 12 and 25 copies of each IS element were found in chromosomal DNAs by Southern analyses. While characteristic fingerprint patterns created by IS element-specific probes were observed with genomic DNA of different S. solfataricus strains, no homologous sequences were identified in DNA of other well-characterized strains of the order Sulfolobales.
Figures





Similar articles
-
An insertion element of the extremely thermophilic archaeon Sulfolobus solfataricus transposes into the endogenous beta-galactosidase gene.Mol Gen Genet. 1994 Apr;243(1):91-6. doi: 10.1007/BF00283880. Mol Gen Genet. 1994. PMID: 8190076
-
Gamma Ray-induced Mutations in pyrEF Genes in Frankia casuarinae Strain CcI3.Microbes Environ. 2025;40(1):ME24062. doi: 10.1264/jsme2.ME24062. Microbes Environ. 2025. PMID: 40074348 Free PMC article.
-
New insertion sequences of Sulfolobus: functional properties and implications for genome evolution in hyperthermophilic archaea.Mol Microbiol. 2005 Jan;55(1):312-25. doi: 10.1111/j.1365-2958.2004.04391.x. Mol Microbiol. 2005. PMID: 15612937
-
Shuffling of Sulfolobus genomes by autonomous and non-autonomous mobile elements.Biochem Soc Trans. 2004 Apr;32(Pt 2):179-83. doi: 10.1042/bst0320179. Biochem Soc Trans. 2004. PMID: 15046567 Review.
-
[Hereditary orotic aciduria].Ryoikibetsu Shokogun Shirizu. 2000;(32):343-5. Ryoikibetsu Shokogun Shirizu. 2000. PMID: 11212739 Review. Japanese. No abstract available.
Cited by
-
Genome sequencing of a genetically tractable Pyrococcus furiosus strain reveals a highly dynamic genome.J Bacteriol. 2012 Aug;194(15):4097-106. doi: 10.1128/JB.00439-12. Epub 2012 May 25. J Bacteriol. 2012. PMID: 22636780 Free PMC article.
-
Genome dynamics in a natural archaeal population.Proc Natl Acad Sci U S A. 2007 Feb 6;104(6):1883-8. doi: 10.1073/pnas.0604851104. Epub 2007 Jan 31. Proc Natl Acad Sci U S A. 2007. PMID: 17267615 Free PMC article.
-
pING family of conjugative plasmids from the extremely thermophilic archaeon Sulfolobus islandicus: insights into recombination and conjugation in Crenarchaeota.J Bacteriol. 2000 Dec;182(24):7014-20. doi: 10.1128/JB.182.24.7014-7020.2000. J Bacteriol. 2000. PMID: 11092863 Free PMC article.
-
Molecular characteristics of spontaneous deletions in the hyperthermophilic archaeon Sulfolobus acidocaldarius.J Bacteriol. 2003 Feb;185(4):1266-72. doi: 10.1128/JB.185.4.1266-1272.2003. J Bacteriol. 2003. PMID: 12562797 Free PMC article.
-
Small multicopy, non-integrative shuttle vectors based on the plasmid pRN1 for Sulfolobus acidocaldarius and Sulfolobus solfataricus, model organisms of the (cren-)archaea.Nucleic Acids Res. 2007;35(12):e88. doi: 10.1093/nar/gkm449. Epub 2007 Jun 18. Nucleic Acids Res. 2007. PMID: 17576673 Free PMC article.
References
-
- Aagaard C, Leviev I, Aravalli R N, Forterre P, Prieur D, Garrett G A. General vectors for archaeal hyperthermophiles: strategies based on a mobile intron and a plasmid. FEMS Microbiol Rev. 1996;18:93–104. - PubMed
-
- Ammendola S, Politi L, Scandurra R. Cloning and sequencing of ISC1041 from the archaeon Sulfolobus solfataricus MT-4, a new member of the IS30 family of insertion elements. FEBS Lett. 1998;428:217–223. - PubMed
-
- Aravalli R N, Garrett R A. Shuttle vectors for hyperthermophilic Archaea. Extremophiles. 1997;1:183–191. - PubMed
-
- Bult C J, White O, Olsen G J, Zhou L, Fleischmann R D, et al. Complete genome sequence of the methanogenic archaeon Methanococcus jannaschii. Science. 1996;273:1058–1073. - PubMed
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

