Ro ribonucleoproteins contribute to the resistance of Deinococcus radiodurans to ultraviolet irradiation
- PMID: 10766734
- PMCID: PMC316496
Ro ribonucleoproteins contribute to the resistance of Deinococcus radiodurans to ultraviolet irradiation
Abstract
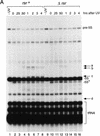
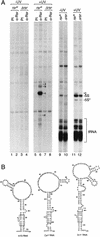
The genome of the radiation-resistant eubacterium Deinococcus radiodurans contains an ortholog of an RNA-binding protein known as the Ro 60-kD autoantigen. This protein, which was previously identified only in higher eukaryotes, is normally bound to small RNAs known as Y RNAs. We show that the Ro protein ortholog Rsr contributes to the resistance of D. radiodurans to UV irradiation. Rsr binds several small RNAs, encoded upstream of rsr, that accumulate following UV irradiation. One of these RNAs resembles a Y RNA. These results suggest that Ro RNPs could similarly contribute to the recovery of higher cells following UV irradiation.
Figures

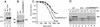

Similar articles
-
Emerging roles for the Ro 60-kDa autoantigen in noncoding RNA metabolism.Wiley Interdiscip Rev RNA. 2011 Sep-Oct;2(5):686-99. doi: 10.1002/wrna.85. Epub 2011 Apr 21. Wiley Interdiscip Rev RNA. 2011. PMID: 21823229 Free PMC article. Review.
-
An ortholog of the Ro autoantigen functions in 23S rRNA maturation in D. radiodurans.Genes Dev. 2007 Jun 1;21(11):1328-39. doi: 10.1101/gad.1548207. Epub 2007 May 17. Genes Dev. 2007. PMID: 17510283 Free PMC article.
-
A lupus-like syndrome develops in mice lacking the Ro 60-kDa protein, a major lupus autoantigen.Proc Natl Acad Sci U S A. 2003 Jun 24;100(13):7503-8. doi: 10.1073/pnas.0832411100. Epub 2003 Jun 3. Proc Natl Acad Sci U S A. 2003. PMID: 12788971 Free PMC article.
-
Crystal structure of Rsr, an ortholog of the antigenic Ro protein, links conformational flexibility to RNA binding activity.J Biol Chem. 2007 May 18;282(20):14960-7. doi: 10.1074/jbc.M611163200. Epub 2007 Mar 28. J Biol Chem. 2007. PMID: 17392270
-
The Ro 60 kDa autoantigen: insights into cellular function and role in autoimmunity.J Mol Med (Berl). 2004 Apr;82(4):232-9. doi: 10.1007/s00109-004-0529-0. J Mol Med (Berl). 2004. PMID: 15168680 Review.
Cited by
-
Sjögren's syndrome--study of autoantigens and autoantibodies.Clin Rev Allergy Immunol. 2007 Jun;32(3):238-51. doi: 10.1007/s12016-007-8003-8. Clin Rev Allergy Immunol. 2007. PMID: 17992591 Review.
-
RNA chaperone activity of protein components of human Ro RNPs.RNA. 2005 Jul;11(7):1084-94. doi: 10.1261/rna.7263905. Epub 2005 May 31. RNA. 2005. PMID: 15928345 Free PMC article.
-
RNA damage in biological conflicts and the diversity of responding RNA repair systems.Nucleic Acids Res. 2016 Oct 14;44(18):8525-8555. doi: 10.1093/nar/gkw722. Epub 2016 Aug 17. Nucleic Acids Res. 2016. PMID: 27536007 Free PMC article. Review.
-
Battle against RNA oxidation: molecular mechanisms for reducing oxidized RNA to protect cells.Wiley Interdiscip Rev RNA. 2014 May-Jun;5(3):335-46. doi: 10.1002/wrna.1214. Epub 2013 Dec 16. Wiley Interdiscip Rev RNA. 2014. PMID: 24375979 Free PMC article. Review.
-
Gut Microbiome and Metabolites in Systemic Lupus Erythematosus: Link, Mechanisms and Intervention.Front Immunol. 2021 Jul 15;12:686501. doi: 10.3389/fimmu.2021.686501. eCollection 2021. Front Immunol. 2021. PMID: 34335588 Free PMC article. Review.
References
-
- Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Struhl K. Current protocols in molecular biology. New York, NY: John Wiley & Sons; 1998.
-
- Battista JR. Against all odds: The survival strategies of Deinococcus radiodurans. Annu Rev Microbiol. 1997;51:203–224. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
