Fission yeast Eso1p is required for establishing sister chromatid cohesion during S phase
- PMID: 10779336
- PMCID: PMC85639
- DOI: 10.1128/MCB.20.10.3459-3469.2000
Fission yeast Eso1p is required for establishing sister chromatid cohesion during S phase
Abstract
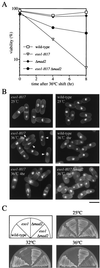
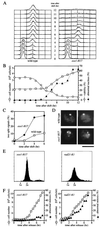
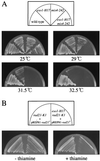
Sister chromatid cohesion is essential for cell viability. We have isolated a novel temperature-sensitive lethal mutant named eso1-H17 that displays spindle assembly checkpoint-dependent mitotic delay and abnormal chromosome segregation. At the permissive temperature, the eso1-H17 mutant shows mild sensitivity to UV irradiation and DNA-damaging chemicals. At the nonpermissive temperature, the mutant is arrested in M phase with a viability loss due to a failure to establish sister chromatid cohesion during S phase. The lethal M-phase arrest phenotype, however, is suppressed by inactivation of a spindle checkpoint. The eso1(+) gene is not essential for the onset and progression of DNA replication but has remarkable genetic interactions with those genes regulating the G(1)-S transition and DNA replication. The N-terminal two-thirds of Eso1p is highly homologous to DNA polymerase eta of budding yeast and humans, and the C-terminal one-third is homologous to budding yeast Eco1p (also called Ctf7p), which is required for the establishment of sister chromatid cohesion. Deletion analysis and determination of the mutation site reveal that the function of the Eco1p/Ctf7p-homologous domain is necessary and sufficient for sister chromatid cohesion. On the other hand, deletion of the DNA polymerase eta domain in Eso1p increases sensitivity to UV irradiation. These results indicate that Eso1p plays a dual role during DNA replication. The C-terminal region acts to establish sister chromatid cohesion, and the N-terminal region presumably catalyzes translesion DNA synthesis when template DNA contains lesions that block regular DNA replication.
Figures






References
-
- Alfa C, Fantes P, Hyams J, McLeod M, Warbrick E. Experiments with fission yeast. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory Press; 1993.
-
- Birkenbihl R P, Subramani S. The rad21 gene product of Schizosaccharomyces pombe is a nuclear, cell cycle-regulated phosphoprotein. J Biol Chem. 1995;270:7703–7711. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
