Conserved plant genes with similarity to mammalian de novo DNA methyltransferases
- PMID: 10781108
- PMCID: PMC18343
- DOI: 10.1073/pnas.97.9.4979
Conserved plant genes with similarity to mammalian de novo DNA methyltransferases
Abstract
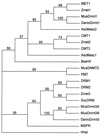
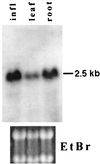
DNA methylation plays a critical role in controlling states of gene activity in most eukaryotic organisms, and it is essential for proper growth and development. Patterns of methylation are established by de novo methyltransferases and maintained by maintenance methyltransferase activities. The Dnmt3 family of de novo DNA methyltransferases has recently been characterized in animals. Here we describe DNA methyltransferase genes from both Arabidopsis and maize that show a high level of sequence similarity to Dnmt3, suggesting that they encode plant de novo methyltransferases. Relative to all known eukaryotic methyltransferases, these plant proteins contain a novel arrangement of the motifs required for DNA methyltransferase catalytic activity. The N termini of these methyltransferases contain a series of ubiquitin-associated (UBA) domains. UBA domains are found in several ubiquitin pathway proteins and in DNA repair enzymes such as Rad23, and they may be involved in ubiquitin binding. The presence of UBA domains provides a possible link between DNA methylation and ubiquitin/proteasome pathways.
Figures



References
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

