Molecular determinants of differential ligand sensitivities of insect ecdysteroid receptors
- PMID: 10805730
- PMCID: PMC85723
- DOI: 10.1128/MCB.20.11.3870-3879.2000
Molecular determinants of differential ligand sensitivities of insect ecdysteroid receptors
Abstract
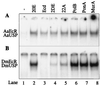
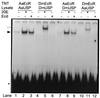
The functional receptor for insect ecdysteroid hormones is a heterodimer consisting of two nuclear hormone receptors, ecdysteroid receptor (EcR) and the retinoid X receptor homologue Ultraspiracle (USP). Although ecdysone is commonly thought to be a hormone precursor and 20-hydroxyecdysone (20E), the physiologically active steroid, little is known about the relative activity of ecdysteroids in various arthropods. As a step toward characterization of potential differential ligand recognition, we have analyzed the activities of various ecdysteroids using gel mobility shift assays and transfection assays in Schneider-2 (S2) cells. Ecdysone showed little activation of the Drosophila melanogaster receptor complex (DmEcR-USP). In contrast, this steroid functioned as a potent ligand for the mosquito Aedes aegypti receptor complex (AaEcR-USP), significantly enhancing DNA binding and transactivating a reporter gene in S2 cells. The mosquito receptor also displayed higher hormone-independent DNA binding activity than the Drosophila receptor. Subunit-swapping experiments indicated that the EcR protein, not the USP protein, was responsible for ligand specificity. Using domain-swapping techniques, we made a series of Aedes and Drosophila EcR chimeric constructs. Differential ligand responsiveness was mapped near the C terminus of the ligand binding domain, within the identity box previously implicated in the dimerization specificity of nuclear receptors. This region includes helices 9 and 10, as determined by comparison with available crystal structures obtained from other nuclear receptors. Site-directed mutagenesis revealed that Phe529 in Aedes EcR, corresponding to Tyr611 in Drosophila EcR, was most critical for ligand specificity and hormone-independent DNA binding activity. These results demonstrated that ecdysone could function as a bona fide ligand in a species-specific manner.
Figures
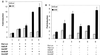




Similar articles
-
Requirement of co-factors for the ligand-mediated activity of the insect ecdysteroid receptor in yeast.J Mol Endocrinol. 2001 Oct;27(2):191-209. doi: 10.1677/jme.0.0270191. J Mol Endocrinol. 2001. PMID: 11564603
-
DNA binding and transactivation characteristics of the mosquito ecdysone receptor-Ultraspiracle complex.J Biol Chem. 1998 Oct 16;273(42):27531-40. doi: 10.1074/jbc.273.42.27531. J Biol Chem. 1998. PMID: 9765285
-
Functional ecdysone receptor is the product of EcR and Ultraspiracle genes.Nature. 1993 Dec 2;366(6454):476-9. doi: 10.1038/366476a0. Nature. 1993. PMID: 8247157
-
Ecdysone receptors: from the Ashburner model to structural biology.Annu Rev Entomol. 2013;58:251-71. doi: 10.1146/annurev-ento-120811-153610. Epub 2012 Oct 9. Annu Rev Entomol. 2013. PMID: 23072463 Review.
-
Arthropod nuclear receptors and their role in molting.FEBS J. 2009 Nov;276(21):6128-57. doi: 10.1111/j.1742-4658.2009.07347.x. Epub 2009 Sep 30. FEBS J. 2009. PMID: 19796154 Review.
Cited by
-
Evaluation of ecdysteroid antisera for a competitive enzyme immunoassay and extraction procedures for the measurement of mosquito ecdysteroids.Gen Comp Endocrinol. 2017 Nov 1;253:60-69. doi: 10.1016/j.ygcen.2017.08.028. Epub 2017 Sep 1. Gen Comp Endocrinol. 2017. PMID: 28866256 Free PMC article.
-
Practical uses for ecdysteroids in mammals including humans: an update.J Insect Sci. 2003;3:7. doi: 10.1093/jis/3.1.7. J Insect Sci. 2003. PMID: 15844229 Free PMC article. Review.
-
Juvenile hormone-regulated alternative splicing of the taiman gene primes the ecdysteroid response in adult mosquitoes.Proc Natl Acad Sci U S A. 2018 Aug 14;115(33):E7738-E7747. doi: 10.1073/pnas.1808146115. Epub 2018 Jul 30. Proc Natl Acad Sci U S A. 2018. PMID: 30061397 Free PMC article.
-
Reporter gene assays for screening and identification of novel molting hormone- and juvenile hormone-like chemicals.J Pestic Sci. 2021 Feb 20;46(1):29-42. doi: 10.1584/jpestics.D20-079. J Pestic Sci. 2021. PMID: 33746544 Free PMC article.
-
Specific transcriptional responses to juvenile hormone and ecdysone in Drosophila.Insect Biochem Mol Biol. 2007 Jun;37(6):570-8. doi: 10.1016/j.ibmb.2007.03.001. Epub 2007 Mar 12. Insect Biochem Mol Biol. 2007. PMID: 17517334 Free PMC article.
References
-
- Antoniewski C, O'Grady M S, Edmondson R G, Lassieur S M, Benes H. Characterization of an EcR/USP heterodimer target site that mediates ecdysone responsiveness of the Drosophila Lsp-2 gene. Mol Gen Genet. 1995;249:545–556. - PubMed
-
- Borovsky D, Whisenton L R, Thomas B R, Fuchs M S. Biosynthesis and distribution of ecdysone and 20-OH-ecdysone in Aedes-aegypti. Arch Insect Biochem Physiol. 1986;3:19–30.
-
- Bourguet W, Ruff M, Chambon P, Gronemeyer H, Moras D. Crystal structure of the ligand-binding domain of the human nuclear receptor RXR-alpha. Nature. 1995;375:377–382. - PubMed
-
- Bownes M. Expression of the genes coding for vitellogenin (yolk protein) Annu Rev Entomol. 1986;31:507–531.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
