Binding of bisubstrate analog promotes large structural changes in the unregulated catalytic trimer of aspartate transcarbamoylase: implications for allosteric regulation
- PMID: 10805770
- PMCID: PMC25784
- DOI: 10.1073/pnas.090087197
Binding of bisubstrate analog promotes large structural changes in the unregulated catalytic trimer of aspartate transcarbamoylase: implications for allosteric regulation
Abstract
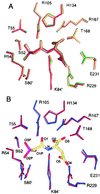
A central problem in understanding enzyme regulation is to define the conformational states that account for allosteric changes in catalytic activity. For Escherichia coli aspartate transcarbamoylase (ATCase; EC) the active, relaxed (R state) holoenzyme is generally assumed to be represented by the crystal structure of the complex of the holoenzyme with the bisubstrate analog N-phosphonacetyl-L-aspartate (PALA). It is unclear, however, which conformational differences between the unliganded, inactive, taut (T state) holoenzyme and the PALA complex are attributable to localized effects of inhibitor binding as contrasted to the allosteric transition. To define the conformational changes in the isolated, nonallosteric C trimer resulting from the binding of PALA, we determined the 1.95-A resolution crystal structure of the C trimer-PALA complex. In contrast to the free C trimer, the PALA-bound trimer exhibits approximate threefold symmetry. Conformational changes in the C trimer upon PALA binding include ordering of two active site loops and closure of the hinge relating the N- and C-terminal domains. The C trimer-PALA structure closely resembles the liganded C subunits in the PALA-bound holoenzyme. This similarity suggests that the pronounced hinge closure and other changes promoted by PALA binding to the holoenzyme are stabilized by ligand binding. Consequently, the conformational changes attributable to the allosteric transition of the holoenzyme remain to be defined.
Figures


Similar articles
-
Peptide-protein interaction markedly alters the functional properties of the catalytic subunit of aspartate transcarbamoylase.Protein Sci. 1993 Jan;2(1):103-12. doi: 10.1002/pro.5560020111. Protein Sci. 1993. PMID: 8443583 Free PMC article.
-
Assessment of the allosteric mechanism of aspartate transcarbamoylase based on the crystalline structure of the unregulated catalytic subunit.Proc Natl Acad Sci U S A. 1999 May 11;96(10):5388-93. doi: 10.1073/pnas.96.10.5388. Proc Natl Acad Sci U S A. 1999. PMID: 10318893 Free PMC article.
-
Cooperative binding of the bisubstrate analog N-(phosphonacetyl)-L-aspartate to aspartate transcarbamoylase and the heterotropic effects of ATP and CTP.J Biol Chem. 1989 Feb 15;264(5):2476-81. J Biol Chem. 1989. PMID: 2644262
-
Structure and mechanisms of Escherichia coli aspartate transcarbamoylase.Acc Chem Res. 2012 Mar 20;45(3):444-53. doi: 10.1021/ar200166p. Epub 2011 Oct 19. Acc Chem Res. 2012. PMID: 22011033 Free PMC article. Review.
-
Allostery and cooperativity in Escherichia coli aspartate transcarbamoylase.Arch Biochem Biophys. 2012 Mar 15;519(2):81-90. doi: 10.1016/j.abb.2011.10.024. Epub 2011 Dec 16. Arch Biochem Biophys. 2012. PMID: 22198283 Free PMC article. Review.
Cited by
-
Allosteric regulation of catalytic activity: Escherichia coli aspartate transcarbamoylase versus yeast chorismate mutase.Microbiol Mol Biol Rev. 2001 Sep;65(3):404-21, table of contents. doi: 10.1128/MMBR.65.3.404-421.2001. Microbiol Mol Biol Rev. 2001. PMID: 11528003 Free PMC article. Review.
-
Oxamic transcarbamylase of Escherichia coli is encoded by the three genes allFGH (formerly fdrA, ylbE, and ylbF).Appl Environ Microbiol. 2024 Jul 24;90(7):e0095724. doi: 10.1128/aem.00957-24. Epub 2024 Jun 18. Appl Environ Microbiol. 2024. PMID: 38888336 Free PMC article.
-
Ligand-induced changes in the conformational stability and flexibility of glutamate dehydrogenase and their role in catalysis and regulation.Protein Sci. 2010 Oct;19(10):1820-9. doi: 10.1002/pro.459. Protein Sci. 2010. PMID: 20665690 Free PMC article.
-
Automated protein crystal structure determination using ELVES.Proc Natl Acad Sci U S A. 2004 Feb 10;101(6):1537-42. doi: 10.1073/pnas.0306241101. Epub 2004 Jan 29. Proc Natl Acad Sci U S A. 2004. PMID: 14752198 Free PMC article.
-
Structural Basis of Sequential and Concerted Cooperativity.Biomolecules. 2022 Nov 7;12(11):1651. doi: 10.3390/biom12111651. Biomolecules. 2022. PMID: 36359000 Free PMC article.
References
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

