MEI-1/MEI-2 katanin-like microtubule severing activity is required for Caenorhabditis elegans meiosis
- PMID: 10809666
- PMCID: PMC316576
MEI-1/MEI-2 katanin-like microtubule severing activity is required for Caenorhabditis elegans meiosis
Abstract
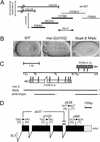
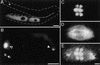
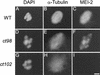
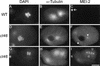
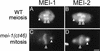
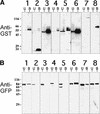
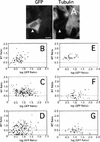
The Caenorhabditis elegans meiotic spindle is morphologically distinct from the first mitotic spindle, yet both structures form in the same cytoplasm approximately 20 minutes apart. The mei-1 and mei-2 genes of C. elegans are required for the establishment of the oocyte meiotic spindle but are not required for mitotic spindle function. mei-1 encodes an AAA ATPase family member with similarity to the p60 catalytic subunit of the heterodimeric sea urchin microtubule-severing protein, katanin. We report that mei-2 encodes a 280-amino acid protein containing a region similar to the p80-targeting subunit of katanin. MEI-1 and MEI-2 antibodies decorate the polar ends of meiotic spindle microtubules and meiotic chromatin. We find that the subcellular location of MEI-2 depends on wild-type mei-1 activity and vice versa. These experiments, combined with MEI-1 and MEI-2's similarity to p60 and p80 katanin, suggest that the C. elegans proteins function as a complex. In support of this idea, MEI-1 and MEI-2 physically associate in HeLa cells. Furthermore, co-expression of MEI-1 and MEI-2 in HeLa cells results in the disassembly of microtubules. These data lead us to conclude that MEI-1/MEI-2 microtubule-severing activity is required for meiotic spindle organization in C. elegans.
Figures


Similar articles
-
MEI-1/katanin is required for translocation of the meiosis I spindle to the oocyte cortex in C elegans.Dev Biol. 2003 Aug 1;260(1):245-59. doi: 10.1016/s0012-1606(03)00216-1. Dev Biol. 2003. PMID: 12885567
-
Microtubule-severing activity of the AAA+ ATPase Katanin is essential for female meiotic spindle assembly.Development. 2016 Oct 1;143(19):3604-3614. doi: 10.1242/dev.140830. Epub 2016 Aug 30. Development. 2016. PMID: 27578779
-
Mutations of a redundant alpha-tubulin gene affect Caenorhabditis elegans early embryonic cleavage via MEI-1/katanin-dependent and -independent pathways.Genetics. 2005 May;170(1):115-26. doi: 10.1534/genetics.104.030106. Epub 2005 Mar 21. Genetics. 2005. PMID: 15781712 Free PMC article.
-
The elegans of spindle assembly.Cell Mol Life Sci. 2010 Jul;67(13):2195-213. doi: 10.1007/s00018-010-0324-8. Epub 2010 Mar 26. Cell Mol Life Sci. 2010. PMID: 20339898 Free PMC article. Review.
-
Cellular Samurai: katanin and the severing of microtubules.J Cell Sci. 2000 Aug;113 ( Pt 16):2821-7. doi: 10.1242/jcs.113.16.2821. J Cell Sci. 2000. PMID: 10910766 Review.
Cited by
-
Models, Regulations, and Functions of Microtubule Severing by Katanin.ISRN Mol Biol. 2012 Sep 27;2012:596289. doi: 10.5402/2012/596289. eCollection 2012. ISRN Mol Biol. 2012. PMID: 27335666 Free PMC article. Review.
-
Multiple roles of endocytosis and autophagy in intracellular remodeling during oocyte-to-embryo transition.Proc Jpn Acad Ser B Phys Biol Sci. 2022;98(5):207-221. doi: 10.2183/pjab.98.013. Proc Jpn Acad Ser B Phys Biol Sci. 2022. PMID: 35545527 Free PMC article. Review.
-
Roles for 147 embryonic lethal genes on C.elegans chromosome I identified by RNA interference and video microscopy.EMBO J. 2001 Aug 1;20(15):3984-92. doi: 10.1093/emboj/20.15.3984. EMBO J. 2001. PMID: 11483502 Free PMC article.
-
Asymmetric spindle positioning.Curr Opin Cell Biol. 2006 Feb;18(1):79-85. doi: 10.1016/j.ceb.2005.12.006. Epub 2005 Dec 19. Curr Opin Cell Biol. 2006. PMID: 16361093 Free PMC article. Review.
-
Cytokinesis in bloodstream stage Trypanosoma brucei requires a family of katanins and spastin.PLoS One. 2012;7(1):e30367. doi: 10.1371/journal.pone.0030367. Epub 2012 Jan 18. PLoS One. 2012. PMID: 22279588 Free PMC article.
References
-
- Afshar K, Barton NR, Hawley RS, Goldstein LSB. DNA binding and meiotic chromosomal localization of the Drosophila nod kinesin-like protein. Cell. 1995;81:129–138. - PubMed
-
- Albertson DG. Formation of the first cleavage spindle in nematode embryos. Dev Biol. 1984;101:61–72. - PubMed
-
- Albertson DG, Thomson JN. Segregation of holocentric chromosomes at meiosis in the nematode, Caenorhabditis elegans. Chromosome Research. 1993;1:15–26. - PubMed
-
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403–410. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous
