Comparative genomic sequencing identifies novel tissue-specific enhancers and sequence elements for methylation-sensitive factors implicated in Igf2/H19 imprinting
- PMID: 10810089
- PMCID: PMC310880
- DOI: 10.1101/gr.10.5.664
Comparative genomic sequencing identifies novel tissue-specific enhancers and sequence elements for methylation-sensitive factors implicated in Igf2/H19 imprinting
Abstract
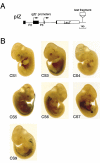
A differentially methylated region (DMR) and endoderm-specific enhancers, located upstream and downstream of the mouse H19 gene, respectively, are known to be essential for the reciprocal imprinting of Igf2 and H19. To explain the same imprinting patterns in non-endodermal tissues, additional enhancers have been hypothesized. We determined and compared the sequences of human and mouse H19 over 40 kb and identified 10 evolutionarily conserved downstream segments, 2 of which were coincident with the known enhancers. Reporter assays in transgenic mice showed that 5 of the other 8 segments functioned as enhancers in specific mesodermal and/or ectodermal tissues. We also identified a conserved 39-bp element that appeared repeatedly within the DMR and formed complexes with specific nuclear factors. Binding of one of the factors was inhibited when the target sequence contained methylated CpGs. These complexes may contribute to the presumed boundary function of the unmethylated DMR, which is proposed to insulate maternal Igf2 from the enhancers. Our results demonstrate that comparative genomic sequencing is highly efficient in identifying regulatory elements.
Figures




References
-
- Ainscough JF-X, Koide T, Tada M, Barton SC, Surani MA. Imprinting of Igf2 and H19 from a 130 kb YAC transgene. Development. 1997;124:3621–3632. - PubMed
-
- Bartolomei MS, Tilghman SM. Parental imprinting of mouse chromosome 7. Semin Dev Biol. 1992;3:107–117.
-
- Bell AC, West AG, Felsenfeld G. The protein CTCF is required for the enhancer blocking activity of vertebrate insulators. Cell. 1999;98:387–396. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous
