Genome evolution of wild barley (Hordeum spontaneum) by BARE-1 retrotransposon dynamics in response to sharp microclimatic divergence
- PMID: 10823912
- PMCID: PMC18673
- DOI: 10.1073/pnas.110587497
Genome evolution of wild barley (Hordeum spontaneum) by BARE-1 retrotransposon dynamics in response to sharp microclimatic divergence
Abstract
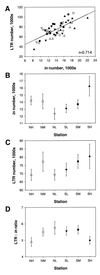
The replicative spread of retrotransposons in the genome creates new insertional polymorphisms, increasing retrotransposon numbers and potentially both their share of the genome and genome size. The BARE-1 retrotransposon constitutes a major, dispersed, active component of Hordeum genomes, and BARE-1 number is positively correlated with genome size. We have examined genome size and BARE-1 insertion patterns and number in wild barley, Hordeum spontaneum, in Evolution Canyon, Lower Nahal Oren, Mount Carmel, Israel, along a transect presenting sharply differing microclimates. BARE-1 has been sufficiently active for its insertional pattern to resolve individuals in a way consonant with their ecogeographical distribution in the canyon and to distinguish them from provenances outside the canyon. On both slopes, but especially on the drier south-facing slope, a simultaneous increase in the BARE-1 copy number and a decrease in the relative number lost through recombination, as measured by the abundance of solo long terminal repeats, appear to have driven the BARE-1 share of the genome upward with the height and dryness of the slope. The lower recombinational loss would favor maintenance of more full-length copies, enhancing the ability of the BARE-1 family to contribute to genome size growth. These local data are consistent with regional trends for BARE-1 in H. spontaneum across Israel and therefore may reflect adaptive selection for increasing genome size through retrotransposon activity.
Figures


Comment in
-
Retrotransposon-mediated genome evolution on a local ecological scale.Proc Natl Acad Sci U S A. 2000 Jun 6;97(12):6250-2. doi: 10.1073/pnas.97.12.6250. Proc Natl Acad Sci U S A. 2000. PMID: 10841529 Free PMC article. No abstract available.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

