Cyanobacterial sulfide-quinone reductase: cloning and heterologous expression
- PMID: 10852862
- PMCID: PMC101880
- DOI: 10.1128/JB.182.12.3336-3344.2000
Cyanobacterial sulfide-quinone reductase: cloning and heterologous expression
Abstract
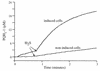
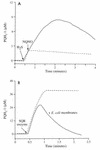
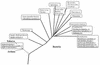
The gene encoding sulfide-quinone reductase (SQR; E.C.1.8.5.'), the enzyme catalyzing the first step of anoxygenic photosynthesis in the filamentous cyanobacterium Oscillatoria limnetica, was cloned by use of amino acid sequences of tryptic peptides as well as sequences conserved in the Rhodobacter capsulatus SQR and in an open reading frame found in the genome of Aquifex aeolicus. SQR activity was also detected in the unicellular cyanobacterium Aphanothece halophytica following sulfide induction, with a V(max) of 180 micromol of plastoquinone-1 (PQ-1) reduced/mg of chlorophyll/h and apparent K(m) values of 20 and 40 microM for sulfide and quinone, respectively. Based on the conserved sequences, the gene encoding A. halophytica SQR was also cloned. The SQR polypeptides deduced from the two cyanobacterial genes consist of 436 amino acids for O. limnetica SQR and 437 amino acids for A. halophytica SQR and show 58% identity and 74% similarity. The calculated molecular mass is about 48 kDa for both proteins; the theoretical isoelectric points are 7.7 and 5.6 and the net charges at a neutral pH are 0 and -14 for O. limnetica SQR and A. halophytica SQR, respectively. A search of databases showed SQR homologs in the genomes of the cyanobacterium Anabaena PCC7120 as well as the chemolithotrophic bacteria Shewanella putrefaciens and Thiobacillus ferrooxidans. All SQR enzymes contain characteristic flavin adenine dinucleotide binding fingerprints. The cyanobacterial proteins were expressed in Escherichia coli under the control of the T7 promoter. Membranes isolated from E. coli cells expressing A. halophytica SQR performed sulfide-dependent PQ-1 reduction that was sensitive to the quinone analog inhibitor 2n-nonyl-4-hydroxyquinoline-N-oxide. The wide distribution of SQR genes emphasizes the important role of SQR in the sulfur cycle in nature.
Figures




Similar articles
-
Sulfide-quinone reductase from Rhodobacter capsulatus. Purification, cloning, and expression.J Biol Chem. 1997 Apr 11;272(15):9890-4. doi: 10.1074/jbc.272.15.9890. J Biol Chem. 1997. PMID: 9092526
-
Purification and characterization of sulfide-quinone reductase, a novel enzyme driving anoxygenic photosynthesis in Oscillatoria limnetica.J Biol Chem. 1994 Feb 25;269(8):5705-11. J Biol Chem. 1994. PMID: 8119908
-
Sulfide-induced sulfide-quinone reductase activity in thylakoids of Oscillatoria limnetica.J Biol Chem. 1991 Jan 5;266(1):104-11. J Biol Chem. 1991. PMID: 1898723
-
Purification and characterization of sulfide:quinone oxidoreductase from an acidophilic iron-oxidizing bacterium, Acidithiobacillus ferrooxidans.Biosci Biotechnol Biochem. 2007 Nov;71(11):2735-42. doi: 10.1271/bbb.70332. Epub 2007 Nov 7. Biosci Biotechnol Biochem. 2007. PMID: 17986789
-
A new structure-based classification of sulfide:quinone oxidoreductases.Proteins. 2010 Apr;78(5):1073-83. doi: 10.1002/prot.22665. Proteins. 2010. PMID: 20077566 Review.
Cited by
-
H2S and reactive sulfur signaling at the host-bacterial pathogen interface.J Biol Chem. 2020 Sep 18;295(38):13150-13168. doi: 10.1074/jbc.REV120.011304. Epub 2020 Jul 22. J Biol Chem. 2020. PMID: 32699012 Free PMC article. Review.
-
Photosynthetic Versatility in the Genome of Geitlerinema sp. PCC 9228 (Formerly Oscillatoria limnetica 'Solar Lake'), a Model Anoxygenic Photosynthetic Cyanobacterium.Front Microbiol. 2016 Oct 13;7:1546. doi: 10.3389/fmicb.2016.01546. eCollection 2016. Front Microbiol. 2016. PMID: 27790189 Free PMC article.
-
Sulfide:quinone oxidoreductase from echiuran worm Urechis unicinctus.Mar Biotechnol (NY). 2011 Feb;13(1):93-107. doi: 10.1007/s10126-010-9273-3. Epub 2010 Apr 25. Mar Biotechnol (NY). 2011. PMID: 20419499
-
Exploring the Origins and Evolution of Oxygenic and Anoxygenic Photosynthesis in Deeply Branched Cyanobacteriota.Mol Biol Evol. 2024 Aug 2;41(8):msae151. doi: 10.1093/molbev/msae151. Mol Biol Evol. 2024. PMID: 39041196 Free PMC article.
-
Unraveling the Physiological Roles of the Cyanobacterium Geitlerinema sp. BBD and Other Black Band Disease Community Members through Genomic Analysis of a Mixed Culture.PLoS One. 2016 Jun 23;11(6):e0157953. doi: 10.1371/journal.pone.0157953. eCollection 2016. PLoS One. 2016. PMID: 27336619 Free PMC article.
References
-
- Arieli B, Padan E, Shahak Y. Sulfide-induced sulfide-quinone reductase activity in thylakoids of Oscillatoria limnetica. J Biol Chem. 1991;266:104–111. - PubMed
-
- Arieli B, Shahak Y, Taglicht D, Hauska G, Padan E. Purification and characterization of sulfide-quinone reductase, a novel enzyme driving anoxygenic photosynthesis in Oscillatoria limnetica. J Biol Chem. 1994;269:5705–5711. - PubMed
-
- Belkin S, Shahak Y, Padan E. Anoxygenic photosynthetic electron transport. Methods Enzymol. 1988;167:380–386.
-
- Belkin S, Arieli B, Padan E. Sulfide-dependent electron transport in Oscillatoria limnetica. Isr J Bot. 1982;31:199–200.
-
- Belkin S, Padan E. Sulfide-dependent hydrogen evolution in the cyanobacterium Oscillatoria limnetica. FEBS Lett. 1978;94:291–294.
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

