leafy hull sterile1 is a homeotic mutation in a rice MADS box gene affecting rice flower development
- PMID: 10852934
- PMCID: PMC149090
- DOI: 10.1105/tpc.12.6.871
leafy hull sterile1 is a homeotic mutation in a rice MADS box gene affecting rice flower development
Abstract
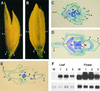
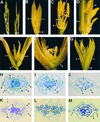
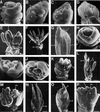
Rice contains several MADS box genes. It has been demonstrated previously that one of these genes, OsMADS1 (for Oryza sativa MADS box gene1), is expressed preferentially in flowers and causes early flowering when ectopically expressed in tobacco plants. In this study, we demonstrated that ectopic expression of OsMADS1 in rice also results in early flowering. To further investigate the role of OsMADS1 during rice flower development, we generated transgenic rice plants expressing altered OsMADS1 genes that contain missense mutations in the MADS domain. There was no visible alteration in the transgenic plants during the vegetative stage. However, transgenic panicles typically exhibited phenotypic alterations, including spikelets consisting of elongated leafy paleae and lemmas that exhibit a feature of open hull, two pairs of leafy palea-like and lemma-like lodicules, a decrease in stamen number, and an increase in the number of carpels. In addition, some spikelets generated an additional floret from the same rachilla. These characteristics are very similar to those of leafy hull sterile1 (lhs1). The map position of OsMADS1 is closely linked to that of lhs1 on chromosome 3. Examination of lhs1 revealed that it contains two missense mutations in the OsMADS1 MADS domain. A genetic complementation experiment showed that the 11.9-kb genomic DNA fragment containing the wild-type OsMADS1 gene rescued the mutant phenotypes. In addition, ectopic expression of the OsMADS1 gene isolated from the lhs1 line resulted in lhs1-conferred phenotypes. These lines of evidence demonstrate that OsMADS1 is the lhs1 gene.
Figures




References
-
- Angenent, G.C., Franken, J., Busscher, M., Colombo, L., and van Tunen, A.J. (1993). Petal and stamen formation in petunia is regulated by the homeotic gene fbp1. Plant J. 4, 101–112. - PubMed
-
- Angenent, G.C., Franken, J., Busscher, M., Weiss, D., and van Tunen, A.J. (1994). Co-suppression of the petunia homeotic gene fbp2 affects the identity of the generative meristem. Plant J. 5, 33–44. - PubMed
-
- Bonhomme, F., Sommer, H., Bernier, G., and Jacqmard, A. (1997). Characterization of SaMADS D from Sinapis alba suggests a dual function of the gene in inflorescence development and floral organogenesis. Plant Mol. Biol. 34, 573–582. - PubMed
-
- Bradley, D., Carpenter, R., Sommer, H., Hartley, N., and Coen, E. (1993). Complementary floral homeotic phenotypes result from opposite orientations of a transposon at the plena locus of Antirrhinum. Cell 72, 85–95. - PubMed
-
- Cacharrón, J., Saedler, H., and Theissen, G. (1999). Expression of MADS box genes ZMM8 and ZMM14 during inflorescence development of Zea mays discriminates between the upper and the lower floret of each spikelet. Dev. Genes Evol. 209, 411–420. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

