The bases of crown gall tumorigenesis
- PMID: 10869063
- PMCID: PMC94570
- DOI: 10.1128/JB.182.14.3885-3895.2000
The bases of crown gall tumorigenesis
Figures

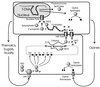
References
-
- Alt-Mörbe J, Stryker J L, Fuqua C, Li P-L, Farrand S K, Winans S C. The conjugal transfer system of Agrobacterium tumefaciens octopine-type Ti plasmids is closely related to the transfer system of an IncP plasmid and distantly related to Ti plasmid vir genes. J Bacteriol. 1996;178:4248–4257. - PMC - PubMed
-
- Barker R F, Idler K B, Thompson D V, Kemp J D. Nucleotide sequence of the T-DNA region from the Agrobacterium tumefaciens octopine Ti plasmid pTi15955. Plant Mol Biol. 1983;2:335–350. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

