CHRK1, a chitinase-related receptor-like kinase in tobacco
- PMID: 10889239
- PMCID: PMC59053
- DOI: 10.1104/pp.123.3.905
CHRK1, a chitinase-related receptor-like kinase in tobacco
Abstract
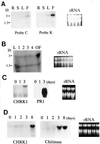
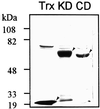
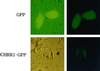
A cDNA encoding a chitinase-related receptor-like kinase, designated CHRK1, was isolated from tobacco (Nicotiana tabacum). The C-terminal kinase domain (KD) of CHRK1 contained all of the conserved amino acids of serine/threonine protein kinases. The putative extracellular domain was closely related to the class V chitinase of tobacco and to microbial chitinases. CHRK1 mRNA accumulation was strongly stimulated by infection with fungal pathogen and tobacco mosaic virus. Amino acid-sequence analysis revealed that the chitinase-like domain of CHRK1 lacked the essential glutamic acid residue required for chitinase activity. The recombinant chitinase-like domain did not show any catalytic activity for either oligomeric or polymeric chitin substrates. The recombinant KD of CHRK1 exhibited autophosphorylation, but the mutant KD with a mutation in the essential ATP-binding site did not, suggesting that CHRK1 encoded a functional kinase. CHRK1 was detected in membrane fractions of tobacco BY2 cells. Furthermore, CHRK1-GFP fusion protein was localized in plasma membranes when it was expressed in animal cells. This is the first report of a new type of receptor-like kinase containing a chitinase-like sequence in the putative extracellular domain.
Figures





References
-
- Baureithel K, Felix G, Boller T. Specific, high affinity binding of chitin fragments to tomato cells and membranes: competitive inhibition of binding by derivatives of chitooligosaccharides and a Nod factor of Rhizobium. J Biol Chem. 1994;269:17931–17938. - PubMed
-
- Becraft PW, Stinard PS, McCarty DR. CRINKLY4: a TNFR-like receptor kinase involved in maize epidermal differentiation. Science. 1996;273:1406–1409. - PubMed
-
- Benhamou N. Elicitor-induced plant defense pathways. Trends Plant Sci. 1996;1:233–240.
-
- Bollag DM, Rozycki MD, Edelstein SJ. Protein Methods. New York: Wiley-Liss; 1996.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

