Modulation of histone acetyltransferase activity through interaction of epstein-barr nuclear antigen 3C with prothymosin alpha
- PMID: 10891508
- PMCID: PMC86050
- DOI: 10.1128/MCB.20.15.5722-5735.2000
Modulation of histone acetyltransferase activity through interaction of epstein-barr nuclear antigen 3C with prothymosin alpha
Abstract
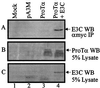
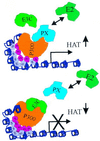
The Epstein-Barr virus (EBV) nuclear antigen 3C (EBNA3C) is essential for EBV-dependent immortalization of human primary B lymphocytes. Genetic analysis indicated that amino acids 365 to 992 are important for EBV-mediated immortalization of B lymphocytes. We demonstrate that this region of EBNA3C critical for immortalization interacts with prothymosin alpha (ProTalpha), a cellular protein previously identified to be important for cell division and proliferation. This interaction maps to a region downstream of amino acid 365 known to be involved in transcription regulation and critical for EBV-mediated transformation of primary B lymphocytes. Additionally, we show that EBNA3C also interacts with p300, a cellular acetyltransferase. This interaction suggests a possible role in regulation of histone acetylation and chromatin remodeling. An increase in histone acetylation was observed in EBV-transformed lymphoblastoid cell lines, which is consistent with increased cellular gene expression. These cells express the entire repertoire of latent nuclear antigens, including EBNA3C. Expression of EBNA3C in cells with increased acetyltransferase activity mediated by the EBV transactivator EBNA2 results in down-modulation of this activity in a dose-responsive manner. The interactions of EBNA3C with ProTalpha and p300 provide new evidence implicating this essential EBV protein EBNA3C in modulating the acetylation of cellular factors, including histones. Hence, EBNA3C plays a critical role in balancing cellular transcriptional events by linking the biological property of mediating inhibition of EBNA2 transcription activation and the observed histone acetyltransferase activity, thereby orchestrating immortalization of EBV-infected cells.
Figures










References
-
- Alfieri C, Birkenbach M, Kieff E. Early events in Epstein-Barr virus infection of human B lymphocytes. Virology. 1991;181:595–608. . (Erratum, 185:946.) - PubMed
-
- Ambinder R F. Human lymphotropic viruses associated with lymphoid malignancy: Epstein-Barr and HTLV-1. Hematol Oncol Clin N Am. 1990;4:821–833. - PubMed
-
- Artavanis-Tsakonas S, Matsuno K, Fortini M E. Notch signaling. Science. 1995;268:225–232. - PubMed
-
- Artavanis-Tsakonas S, Rand M D, Lake R J. Notch signaling: cell fate control and signal integration in development. Science. 1999;284:770–776. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous
