Myotubularin, a protein tyrosine phosphatase mutated in myotubular myopathy, dephosphorylates the lipid second messenger, phosphatidylinositol 3-phosphate
- PMID: 10900271
- PMCID: PMC16795
- DOI: 10.1073/pnas.160255697
Myotubularin, a protein tyrosine phosphatase mutated in myotubular myopathy, dephosphorylates the lipid second messenger, phosphatidylinositol 3-phosphate
Abstract
The lipid second messenger phosphatidylinositol 3-phosphate [PI(3)P] plays a crucial role in intracellular membrane trafficking. We report here that myotubularin, a protein tyrosine phosphatase required for muscle cell differentiation, is a potent PI(3)P phosphatase. Recombinant human myotubularin specifically dephosphorylates PI(3)P in vitro. Overexpression of a catalytically inactive substrate-trapping myotubularin mutant (C375S) in human 293 cells increases PI(3)P levels relative to that of cells overexpressing the wild-type enzyme, demonstrating that PI(3)P is a substrate for myotubularin in vivo. In addition, a Saccharomyces cerevisiae strain in which the myotubularin-like gene (YJR110w) is disrupted also exhibits increased PI(3)P levels. Both the recombinant yeast enzyme and a human myotubularin-related protein (KIAA0371) are able to dephosphorylate PI(3)P in vitro, suggesting that this activity is intrinsic to all myotubularin family members. Mutations in the MTM1 gene that cause human myotubular myopathy dramatically reduce the ability of the phosphatase to dephosphorylate PI(3)P. Our findings provide evidence that myotubularin exerts its effects during myogenesis by regulating cellular levels of the inositol lipid PI(3)P.
Figures




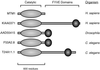
References
-
- Spiro A J, Shy G M, Gonatas N K. Arch Neurol. 1966;14:1–14. - PubMed
-
- Van Wijngaarden G K, Fleury P, Bethlem J, Meijer A E F H. Neurology. 1969;19:901–908. - PubMed
-
- Sarnat H B, Roth S I, Jimenez J F. Can J Neurol Sci. 1981;8:313–320. - PubMed
-
- Laporte J, Hu L J, Kretz C, Mandel J-L, Kioschis P, Coy J F, Klauck S M, Poutska A, Dahl N. Nat Genet. 1996;13:175–182. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

