Dmc1 of Schizosaccharomyces pombe plays a role in meiotic recombination
- PMID: 10908327
- PMCID: PMC102652
- DOI: 10.1093/nar/28.14.2709
Dmc1 of Schizosaccharomyces pombe plays a role in meiotic recombination
Abstract
We report here a Schizosaccharomyces pombe gene (dmc1(+)) that resembles budding yeast DMC1 in the region immediately upstream of the rad24(+) gene. We showed by northern and Southern blot analysis that dmc1(+) and rad24(+) are co-transcribed as a bicistronic mRNA of 2.8 kb with meiotic specificity, whereas rad24(+) itself is constitutively transcribed as a 1.0-kb mRNA species during meiosis. Induction of the bicistronic transcript is under the control of a meiosis-specific transcription factor, Ste11. Disruption of both dmc1(+) and rad24(+) had no effect on mitosis or spore formation, and dmc1Delta cells displayed no change in sensitivity to UV or gamma irradiation relative to the wild type. Tetrad analysis indicated that Dmc1 is involved in meiotic recombination. Analysis of gene conversion frequencies using single and double mutants of dmc1 and rhp51 indicated that both Dmc1 and Rhp51 function in meiotic gene conversion. These observations, together with a high level of sequence identity, indicate that the dmc1(+) gene of S. POMBE: is a structural homolog of budding yeast DMC1, sharing both similar and distinct functions in meiosis.
Figures
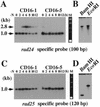

Similar articles
-
Cloning and sequence analysis of rhp51+, a Schizosaccharomyces pombe homolog of the Saccharomyces cerevisiae RAD51 gene.Gene. 1994 May 16;142(2):207-11. doi: 10.1016/0378-1119(94)90262-3. Gene. 1994. PMID: 8194753
-
From meiosis to postmeiotic events: uncovering the molecular roles of the meiosis-specific recombinase Dmc1.FEBS J. 2010 Feb;277(3):590-8. doi: 10.1111/j.1742-4658.2009.07503.x. Epub 2009 Dec 15. FEBS J. 2010. PMID: 20015079 Review.
-
Human and mouse homologs of Schizosaccharomyces pombe rad1(+) and Saccharomyces cerevisiae RAD17: linkage to checkpoint control and mammalian meiosis.Genes Dev. 1998 Aug 15;12(16):2560-73. doi: 10.1101/gad.12.16.2560. Genes Dev. 1998. PMID: 9716408 Free PMC article.
-
Homologous recombination in the fission yeast Schizosaccharomyces pombe: different requirements for the rhp51+, rhp54+ and rad22+ genes.Curr Genet. 1997 Mar;31(3):248-54. doi: 10.1007/s002940050202. Curr Genet. 1997. PMID: 9065388
-
Telomeres in meiotic recombination: the yeast side story.Cell Mol Life Sci. 2007 Jan;64(2):125-30. doi: 10.1007/s00018-006-6464-1. Cell Mol Life Sci. 2007. PMID: 17219024 Free PMC article. Review.
Cited by
-
Meiotic versus mitotic recombination: two different routes for double-strand break repair: the different functions of meiotic versus mitotic DSB repair are reflected in different pathway usage and different outcomes.Bioessays. 2010 Dec;32(12):1058-66. doi: 10.1002/bies.201000087. Epub 2010 Oct 21. Bioessays. 2010. PMID: 20967781 Free PMC article. Review.
-
Structural and functional analyses of the DMC1-M200V polymorphism found in the human population.Nucleic Acids Res. 2008 Jul;36(12):4181-90. doi: 10.1093/nar/gkn362. Epub 2008 Jun 19. Nucleic Acids Res. 2008. PMID: 18566005 Free PMC article.
-
Genetic and cytological characterization of the RecA-homologous proteins Rad51 and Dmc1 of Schizosaccharomyces pombe.Curr Genet. 2004 Jan;44(6):317-28. doi: 10.1007/s00294-003-0439-7. Epub 2003 Aug 29. Curr Genet. 2004. PMID: 12955454
-
Many functions of the meiotic cohesin.Chromosome Res. 2010 Dec;18(8):909-24. doi: 10.1007/s10577-010-9169-0. Epub 2010 Nov 18. Chromosome Res. 2010. PMID: 21086039 Review.
-
Insights into the identification and evolutionary conservation of key genes in the transcriptional circuits of meiosis initiation and commitment in budding yeast.FEBS Open Bio. 2023 Dec;13(12):2290-2305. doi: 10.1002/2211-5463.13728. Epub 2023 Nov 14. FEBS Open Bio. 2023. PMID: 37905308 Free PMC article.
References
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

