Alternatively spliced hBRF variants function at different RNA polymerase III promoters
- PMID: 10921893
- PMCID: PMC306597
- DOI: 10.1093/emboj/19.15.4134
Alternatively spliced hBRF variants function at different RNA polymerase III promoters
Abstract
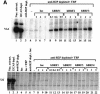
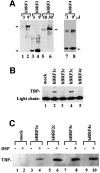
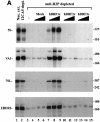
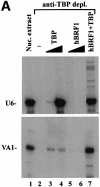
In yeast, a single form of TFIIIB is required for transcription of all RNA polymerase III (pol III) genes. It consists of three subunits: the TATA box-binding protein (TBP), a TFIIB-related factor, BRF, and B". Human TFIIIB is not as well defined and human pol III promoters differ in their requirements for this activity. A human homolog of yeast BRF was shown to be required for transcription at the gene-internal 5S and VA1 promoters. Whether or not it was also involved in transcription from the gene-external human U6 promoter was unclear. We have isolated cDNAs encoding alternatively spliced forms of human BRF that can complex with TBP. Using immunopurified complexes containing the cloned hBRFs, we show that while hBRF1 functions at the 5S, VA1, 7SL and EBER2 promoters, a different variant, hBRF2, is required at the human U6 promoter. Thus, pol III utilizes different TFIIIB complexes at structurally distinct promoters.
Figures









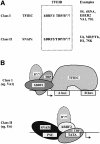
Similar articles
-
Different human TFIIIB activities direct RNA polymerase III transcription from TATA-containing and TATA-less promoters.Genes Dev. 2000 Oct 15;14(20):2650-63. doi: 10.1101/gad.836400. Genes Dev. 2000. PMID: 11040218 Free PMC article.
-
Polymerase (Pol) III TATA box-binding protein (TBP)-associated factor Brf binds to a surface on TBP also required for activated Pol II transcription.Mol Cell Biol. 1998 Mar;18(3):1692-700. doi: 10.1128/MCB.18.3.1692. Mol Cell Biol. 1998. PMID: 9488486 Free PMC article.
-
Structure-function analysis of the human TFIIB-related factor II protein reveals an essential role for the C-terminal domain in RNA polymerase III transcription.Mol Cell Biol. 2005 Nov;25(21):9406-18. doi: 10.1128/MCB.25.21.9406-9418.2005. Mol Cell Biol. 2005. PMID: 16227591 Free PMC article.
-
Comparison of the RNA polymerase III transcription machinery in Schizosaccharomyces pombe, Saccharomyces cerevisiae and human.Nucleic Acids Res. 2001 Jul 1;29(13):2675-90. doi: 10.1093/nar/29.13.2675. Nucleic Acids Res. 2001. PMID: 11433012 Free PMC article. Review.
-
The RNA polymerase III transcription apparatus.J Mol Biol. 2001 Jun 29;310(1):1-26. doi: 10.1006/jmbi.2001.4732. J Mol Biol. 2001. PMID: 11419933 Review. No abstract available.
Cited by
-
Essential roles of Bdp1, a subunit of RNA polymerase III initiation factor TFIIIB, in transcription and tRNA processing.Mol Cell Biol. 2002 May;22(10):3264-75. doi: 10.1128/MCB.22.10.3264-3275.2002. Mol Cell Biol. 2002. PMID: 11971960 Free PMC article.
-
Transcriptional regulation of human small nuclear RNA genes.Biochim Biophys Acta. 2008 May;1779(5):295-305. doi: 10.1016/j.bbagrm.2008.04.001. Epub 2008 Apr 8. Biochim Biophys Acta. 2008. PMID: 18442490 Free PMC article. Review.
-
Different sequence signatures in the upstream regions of plant and animal tRNA genes shape distinct modes of regulation.Nucleic Acids Res. 2011 Apr;39(8):3331-9. doi: 10.1093/nar/gkq1257. Epub 2010 Dec 7. Nucleic Acids Res. 2011. PMID: 21138970 Free PMC article.
-
RNA polymerase III repression by the retinoblastoma tumor suppressor protein.Biochim Biophys Acta. 2013 Mar-Apr;1829(3-4):385-92. doi: 10.1016/j.bbagrm.2012.09.011. Epub 2012 Oct 12. Biochim Biophys Acta. 2013. PMID: 23063750 Free PMC article. Review.
-
The green tea component EGCG inhibits RNA polymerase III transcription.Biochem Biophys Res Commun. 2007 Sep 7;360(4):778-83. doi: 10.1016/j.bbrc.2007.06.114. Epub 2007 Jul 2. Biochem Biophys Res Commun. 2007. PMID: 17624304 Free PMC article.
References
-
- Brow D.A. and Guthrie,C. (1990) Transcription of a yeast U6 snRNA gene requires a polymerase III promoter element in a novel position. Genes Dev., 4, 1345–1356. - PubMed
-
- Buratowski S. and Zhou,H. (1992) A suppressor of TBP mutations encodes an RNA polymerase III transcription factor with homology to TFIIB. Cell, 71, 221–230. - PubMed
-
- Colbert T. and Hahn,S. (1992) A yeast TFIIB-related factor involved in RNA polymerase III transcription. Genes Dev., 6, 1940–1949. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

