Dynamic relocalization of phage phi 29 DNA during replication and the role of the viral protein p16.7
- PMID: 10921898
- PMCID: PMC306615
- DOI: 10.1093/emboj/19.15.4182
Dynamic relocalization of phage phi 29 DNA during replication and the role of the viral protein p16.7
Abstract
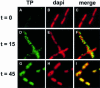
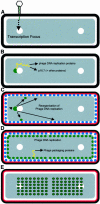
We have examined the localization of DNA replication of the Bacillus subtilis phage phi 29 by immunofluorescence. To determine where phage replication was localized within infected cells, we examined the distribution of phage replication proteins and the sites of incorporation of nucleotide analogues into phage DNA. On initiation of replication, the phage DNA localized to a single focus within the cell, nearly always towards one end of the host cell nucleoid. At later stages of the infection cycle, phage replication was found to have redistributed to multiple sites around the periphery of the nucleoid, just under the cell membrane. Towards the end of the cycle, phage DNA was once again redistributed to become located within the bulk of the nucleoid. Efficient redistribution of replicating phage DNA from the initial replication site to various sites surrounding the nucleoid was found to be dependent on the phage protein p16.7.
Figures



Similar articles
-
The Bacillus subtilis phage phi 29 protein p16.7, involved in phi 29 DNA replication, is a membrane-localized single-stranded DNA-binding protein.J Biol Chem. 2002 Feb 22;277(8):6733-42. doi: 10.1074/jbc.M109312200. Epub 2001 Dec 10. J Biol Chem. 2002. PMID: 11741949
-
Single-stranded DNA binding protein Gp5 of Bacillus subtilis phage Φ29 is required for viral DNA replication in growth-temperature dependent fashion.Biosci Biotechnol Biochem. 2012;76(12):2351-3. doi: 10.1271/bbb.120587. Epub 2012 Dec 7. Biosci Biotechnol Biochem. 2012. PMID: 23221709
-
In vivo functional relationships among terminal proteins of Bacillus subtilis phi 29-related phages.Gene. 1994 Oct 11;148(1):107-12. doi: 10.1016/0378-1119(94)90242-9. Gene. 1994. PMID: 7926823
-
Role of host factors in bacteriophage φ29 DNA replication.Adv Virus Res. 2012;82:351-83. doi: 10.1016/B978-0-12-394621-8.00020-0. Adv Virus Res. 2012. PMID: 22420858 Review.
-
Replication cycle of Bacillus subtilis hydroxymethyluracil-containing phages.Annu Rev Microbiol. 1992;46:95-116. doi: 10.1146/annurev.mi.46.100192.000523. Annu Rev Microbiol. 1992. PMID: 1444272 Review.
Cited by
-
Disclosing the in vivo organization of a viral histone-like protein in Bacillus subtilis mediated by its capacity to recognize the viral genome.Proc Natl Acad Sci U S A. 2012 Apr 10;109(15):5723-8. doi: 10.1073/pnas.1203824109. Epub 2012 Mar 26. Proc Natl Acad Sci U S A. 2012. PMID: 22451942 Free PMC article.
-
Analysis of early promoters of the Bacillus bacteriophage GA-1.J Bacteriol. 2001 Dec;183(23):6965-70. doi: 10.1128/JB.183.23.6965-6970.2001. J Bacteriol. 2001. PMID: 11698389 Free PMC article.
-
The integral membrane protein p16.7 organizes in vivo phi29 DNA replication through interaction with both the terminal protein and ssDNA.EMBO J. 2003 May 1;22(9):2297-306. doi: 10.1093/emboj/cdg221. EMBO J. 2003. PMID: 12727895 Free PMC article.
-
Pleiotropic effect of protein P6 on the viral cycle of bacteriophage phi29.J Bacteriol. 2000 Dec;182(24):6927-32. doi: 10.1128/JB.182.24.6927-6932.2000. J Bacteriol. 2000. PMID: 11092852 Free PMC article.
-
Compartmentalization of phage phi29 DNA replication: interaction between the primer terminal protein and the membrane-associated protein p1.EMBO J. 2000 Oct 16;19(20):5575-84. doi: 10.1093/emboj/19.20.5575. EMBO J. 2000. PMID: 11032825 Free PMC article.
References
-
- Bravo A. and Salas,M. (1997) Initiation of bacteriophage φ29 DNA replication in vivo: assembly of a membrane-associated multiprotein complex. J. Mol. Biol., 269, 102–112. - PubMed
-
- Crucitti P., Lázaro,J.M., Benes,V. and Salas,M. (1998) Bacteriophage φ29 early protein p17 is conditionally required for the first rounds of viral DNA replication. Gene, 223, 135–142. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

