Clinical evaluation of the automated COBAS AMPLICOR HCV MONITOR test version 2.0 for quantifying serum hepatitis C virus RNA and comparison to the quantiplex HCV version 2.0 test
- PMID: 10921954
- PMCID: PMC87152
- DOI: 10.1128/JCM.38.8.2933-2939.2000
Clinical evaluation of the automated COBAS AMPLICOR HCV MONITOR test version 2.0 for quantifying serum hepatitis C virus RNA and comparison to the quantiplex HCV version 2.0 test
Abstract
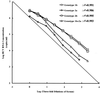
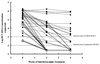
A second-generation hepatitis C virus (HCV) quantitative assay (COBAS AMPLICOR HCV MONITOR Test, version 2.0; COBAS HCM-2) has been developed, with the intention of achieving equivalent quantification of all HCV genotypes and improving assay performance. To evaluate the clinical performance of COBAS HCM-2 and its utility in predicting the response to alpha interferon treatment, sera from 215 chronic hepatitis C patients were analyzed and the results were compared with those obtained by the Quantiplex bDNA HCV RNA, version 2.0, assay (bDNA-2). The COBAS HCM-2 had significantly greater sensitivity than bDNA-2 (94.9 versus 88.4%; P < 0.001) when performed with sera from chronic hepatitis C patients who were viremic by a qualitative PCR test. The standard deviations for the within-run and between-run reproducibilities of COBAS HCM-2 were <0. 1 and <0.2, respectively, and it showed an improved linear range between genotypes with the threefold serial dilutions tested (r(2) = 0.986 to 0.995). The COBAS HCM-2 results were positively correlated with the bDNA-2 results, but the values for COBAS HCM-2 were on average 0.96 log lower than the values for bDNA-2. The mean difference in quantification values between these two assays did not differ among samples with different genotypes (0.70 to 1.00 log). No genotype-dependent difference in viral load was observed. The pretreatment viral load was significantly lower in complete responders. By using multivariate analysis, the viral load 2 weeks after the initiation of alpha interferon treatment was the strongest predictor of a complete response. In conclusion, COBAS HCM-2 demonstrated good sensitivity, linearity, and reproducibility and efficiency equal to that of bDNA-2 for the quantification of HCV genotypes 1 and 2. Hence, this assay provides a rapid and reliable method for the quantification of HCV RNA in serum and is useful for the planning of interferon treatment.
Figures

Similar articles
-
Comparative evaluation of the VERSANT HCV RNA 3.0, QUANTIPLEX HCV RNA 2.0, and COBAS AMPLICOR HCV MONITOR version 2.0 Assays for quantification of hepatitis C virus RNA in serum.J Clin Microbiol. 2002 Feb;40(2):495-500. doi: 10.1128/JCM.40.2.495-500.2002. J Clin Microbiol. 2002. PMID: 11825962 Free PMC article.
-
Clinical application of the Quantiplex HCV RNA 2.0 and Amplicor HCV Monitor assays for quantifying serum hepatitis C virus RNA.J Clin Pathol. 1999 Nov;52(11):807-11. doi: 10.1136/jcp.52.11.807. J Clin Pathol. 1999. PMID: 10690168 Free PMC article.
-
Comparative evaluation of the total hepatitis C virus core antigen, branched-DNA, and amplicor monitor assays in determining viremia for patients with chronic hepatitis C during interferon plus ribavirin combination therapy.J Clin Microbiol. 2003 Jul;41(7):3212-20. doi: 10.1128/JCM.41.7.3212-3220.2003. J Clin Microbiol. 2003. PMID: 12843066 Free PMC article.
-
Assessment of viral loads in patients with chronic hepatitis C with AMPLICOR HCV MONITOR version 1.0, COBAS HCV MONITOR version 2.0, and QUANTIPLEX HCV RNA version 2.0 assays.J Clin Microbiol. 2000 Jul;38(7):2722-5. doi: 10.1128/JCM.38.7.2722-2725.2000. J Clin Microbiol. 2000. PMID: 10878070 Free PMC article.
-
Evaluation of the clinical usefulness of COBAS AMPLICOR HCV MONITOR assay (ver2.0): Comparison with AMPLICOR HCV MONITOR assay (ver1.0) and HCV core protein level.J Med Virol. 2002 Nov;68(3):343-51. doi: 10.1002/jmv.10209. J Med Virol. 2002. PMID: 12226820
Cited by
-
Quantitative assay of hepatitis C virus RNA using an automated extraction system for specific capture with probes and paramagnetic particle separation.J Clin Microbiol. 2003 Feb;41(2):572-5. doi: 10.1128/JCM.41.2.572-575.2003. J Clin Microbiol. 2003. PMID: 12574248 Free PMC article.
-
Comparative evaluation of the VERSANT HCV RNA 3.0, QUANTIPLEX HCV RNA 2.0, and COBAS AMPLICOR HCV MONITOR version 2.0 Assays for quantification of hepatitis C virus RNA in serum.J Clin Microbiol. 2002 Feb;40(2):495-500. doi: 10.1128/JCM.40.2.495-500.2002. J Clin Microbiol. 2002. PMID: 11825962 Free PMC article.
-
Highly sensitive multiplex assay for detection of human immunodeficiency virus type 1 and hepatitis C virus RNA.J Clin Microbiol. 2002 Jul;40(7):2408-19. doi: 10.1128/JCM.40.7.2408-2419.2002. J Clin Microbiol. 2002. PMID: 12089255 Free PMC article.
-
Antinuclear antibody titer and treatment response to peginterferon plus ribavirin for chronic hepatitis C patients.Kaohsiung J Med Sci. 2012 Feb;28(2):86-93. doi: 10.1016/j.kjms.2011.10.031. Epub 2012 Jan 20. Kaohsiung J Med Sci. 2012. PMID: 22313535 Free PMC article. Clinical Trial.
-
The future of viral hepatitis testing: innovations in testing technologies and approaches.BMC Infect Dis. 2017 Nov 1;17(Suppl 1):699. doi: 10.1186/s12879-017-2775-0. BMC Infect Dis. 2017. PMID: 29143676 Free PMC article. Review.
References
-
- Albadalejo J, Alonso R, Antinozzi R, Bogard M, Bourgault A M, Colucci G, Fenner T, Petersen H, Sala E, Vincelette J, Young C. Multicenter evaluation of the COBAS AMPLICOR HCV assay, an integrated PCR system for rapid detection of hepatitis C virus RNA in the diagnostic laboratory. J Clin Microbiol. 1998;36:862–865. - PMC - PubMed
-
- Alter M J, Margolis H S, Krawczynski K, Judson F N, Mares A, Alexander W J, Hu P Y, Miller J K, Gerber M A, Sampliner R E, et al. The natural history of community-acquired hepatitis C in the United States. N Engl J Med. 1992;327:1899–1905. - PubMed
-
- Choo Q L, Kuo G, Weiner A J, Overby L R, Bradley D W, Houghton M. Isolation of a cDNA clone derived from a blood-borne non-A, non-B viral hepatitis genome. Science. 1989;244:359–362. - PubMed
-
- Colucci G, Gutekunst K. Development of a quantitative PCR assay for monitoring HCV viremia levels in patients with chronic hepatitis C. J Viral Hepat. 1997;4:75–78. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

