A hydrogen peroxide-forming NADH oxidase that functions as an alkyl hydroperoxide reductase in Amphibacillus xylanus
- PMID: 10960086
- PMCID: PMC94650
- DOI: 10.1128/JB.182.18.5046-5051.2000
A hydrogen peroxide-forming NADH oxidase that functions as an alkyl hydroperoxide reductase in Amphibacillus xylanus
Abstract
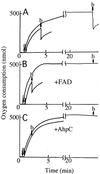
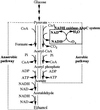
The Amphibacillus xylanus NADH oxidase, which catalyzes the reduction of oxygen to hydrogen peroxide with beta-NADH, can also reduce hydrogen peroxide to water in the presence of free flavin adenine dinucleotide (FAD) or the small disulfide-containing Salmonella enterica AhpC protein. The enzyme has two disulfide bonds, Cys128-Cys131 and Cys337-Cys340, which can act as redox centers in addition to the enzyme-bound FAD (K. Ohnishi, Y. Niimura, M. Hidaka, H. Masaki, H. Suzuki, T. Uozumi, and T. Nishino, J. Biol. Chem. 270:5812-5817, 1995). The NADH-FAD reductase activity was directly dependent on the FAD concentration, with a second-order rate constant of approximately 2.0 x 10(6) M(-1) s(-1). Rapid-reaction studies showed that the reduction of free flavin occurred through enzyme-bound FAD, which was reduced by NADH. The peroxidase activity of NADH oxidase in the presence of FAD resulted from reduction of peroxide by free FADH(2) reduced via enzyme-bound FAD. This peroxidase activity was markedly decreased in the presence of oxygen, since the free FADH(2) is easily oxidized by oxygen, indicating that this enzyme system is unlikely to be functional in aerobic growing cells. The A. xylanus ahpC gene was cloned and overexpressed in Escherichia coli. When the NADH oxidase was coupled with A. xylanus AhpC, the peroxidase activity was not inhibited by oxygen. The V(max) values for hydrogen peroxide and cumene hydroperoxide reduction were both approximately 150 s(-1). The K(m) values for hydrogen peroxide and cumene hydroperoxide were too low to allow accurate determination of their values. Both AhpC and NADH oxidase were induced under aerobic conditions, a clear indication that these proteins are involved in the removal of peroxides under aerobic growing conditions.
Figures


Similar articles
-
Hydrogen peroxide-forming NADH oxidase belonging to the peroxiredoxin oxidoreductase family: existence and physiological role in bacteria.J Bacteriol. 2001 Apr;183(8):2431-8. doi: 10.1128/JB.183.8.2431-2438.2001. J Bacteriol. 2001. PMID: 11274101 Free PMC article.
-
Amphibacillus xylanus NADH oxidase and Salmonella typhimurium alkyl-hydroperoxide reductase flavoprotein components show extremely high scavenging activity for both alkyl hydroperoxide and hydrogen peroxide in the presence of S. typhimurium alkyl-hydroperoxide reductase 22-kDa protein component.J Biol Chem. 1995 Oct 27;270(43):25645-50. doi: 10.1074/jbc.270.43.25645. J Biol Chem. 1995. PMID: 7592740
-
A flavoprotein functional as NADH oxidase from Amphibacillus xylanus Ep01: purification and characterization of the enzyme and structural analysis of its gene.J Bacteriol. 1993 Dec;175(24):7945-50. doi: 10.1128/jb.175.24.7945-7950.1993. J Bacteriol. 1993. PMID: 8253683 Free PMC article.
-
The NADH oxidase-Prx system in Amphibacillus xylanus.Subcell Biochem. 2007;44:195-205. doi: 10.1007/978-1-4020-6051-9_8. Subcell Biochem. 2007. PMID: 18084894 Review.
-
AhpF and other NADH:peroxiredoxin oxidoreductases, homologues of low Mr thioredoxin reductase.Eur J Biochem. 2000 Oct;267(20):6126-33. doi: 10.1046/j.1432-1327.2000.01704.x. Eur J Biochem. 2000. PMID: 11012664 Review.
Cited by
-
NADH oxidase and alkyl hydroperoxide reductase subunit C (peroxiredoxin) from Amphibacillus xylanus form an oligomeric assembly.FEBS Open Bio. 2015 Feb 7;5:124-31. doi: 10.1016/j.fob.2015.01.005. eCollection 2015. FEBS Open Bio. 2015. PMID: 25737838 Free PMC article.
-
H(2)O(2) production in species of the Lactobacillus acidophilus group: a central role for a novel NADH-dependent flavin reductase.Appl Environ Microbiol. 2014 Apr;80(7):2229-39. doi: 10.1128/AEM.04272-13. Epub 2014 Jan 31. Appl Environ Microbiol. 2014. PMID: 24487531 Free PMC article.
-
Hydrogen peroxide-forming NADH oxidase belonging to the peroxiredoxin oxidoreductase family: existence and physiological role in bacteria.J Bacteriol. 2001 Apr;183(8):2431-8. doi: 10.1128/JB.183.8.2431-2438.2001. J Bacteriol. 2001. PMID: 11274101 Free PMC article.
-
Bacterial cyanide oxygenase is a suite of enzymes catalyzing the scavenging and adventitious utilization of cyanide as a nitrogenous growth substrate.J Bacteriol. 2005 Sep;187(18):6396-402. doi: 10.1128/JB.187.18.6396-6402.2005. J Bacteriol. 2005. PMID: 16159773 Free PMC article.
-
Dynamic aspect of reactive oxygen and nitric oxide in oral cavity.J Clin Biochem Nutr. 2008 Jan;42(1):8-13. doi: 10.3164/jcbn.2008002. J Clin Biochem Nutr. 2008. PMID: 18231624 Free PMC article.
References
-
- Alexander N M, Corcoran B J. The reversible dissociation of thyroid iodide peroxidase into apoenzyme and prosthetic group. J Biol Chem. 1962;237:243–248. - PubMed
-
- Beauchamp C, Fridovich I. Superoxide dismutase: improved assays and an assay applicable to acrylamide gels. Anal Biochem. 1971;44:276–287. - PubMed
-
- Bevington . Data reduction and error analysis for the physical sciences. New York, N.Y: McGraw Hill, Inc.; 1969. pp. 235–242.
-
- Castelfranco P, Stumpf P K, Contopouleu R. Fat metabolism in higher plants. J Biol Chem. 1955;214:567–587. - PubMed
-
- Chae H Z, Robison K, Poole L B, Church G, Storz G, Rhee S G. Cloning and sequencing of thiol-specific antioxidant from mammalian brain: alkyl hydroperoxide reductase and thiol-specific antioxidant define a large family of antioxidant enzymes. Proc Natl Acad Sci USA. 1994;91:7017–7021. - PMC - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

