Signal peptide-dependent protein transport in Bacillus subtilis: a genome-based survey of the secretome
- PMID: 10974125
- PMCID: PMC99003
- DOI: 10.1128/MMBR.64.3.515-547.2000
Signal peptide-dependent protein transport in Bacillus subtilis: a genome-based survey of the secretome
Abstract
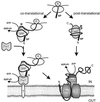
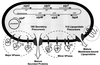
One of the most salient features of Bacillus subtilis and related bacilli is their natural capacity to secrete a variety of proteins into their environment, frequently to high concentrations. This has led to the commercial exploitation of bacilli as major "cell factories" for secreted enzymes. The recent sequencing of the genome of B. subtilis has provided major new impulse for analysis of the molecular mechanisms underlying protein secretion by this organism. Most importantly, the genome sequence has allowed predictions about the composition of the secretome, which includes both the pathways for protein transport and the secreted proteins. The present survey of the secretome describes four distinct pathways for protein export from the cytoplasm and approximately 300 proteins with the potential to be exported. By far the largest number of exported proteins are predicted to follow the major "Sec" pathway for protein secretion. In contrast, the twin-arginine translocation "Tat" pathway, a type IV prepilin-like export pathway for competence development, and ATP-binding cassette transporters can be regarded as "special-purpose" pathways, through which only a few proteins are transported. The properties of distinct classes of amino-terminal signal peptides, directing proteins into the various protein transport pathways, as well as the major components of each pathway are discussed. The predictions and comparisons in this review pinpoint important differences as well as similarities between protein transport systems in B. subtilis and other well-studied organisms, such as Escherichia coli and the yeast Saccharomyces cerevisiae. Thus, they may serve as a lead for future research and applications.
Figures

References
-
- Akita M, Sasaki S, Matsuyama S, Mizushima S. SecA interacts with secretory proteins by recognizing the positive charge at the amino terminus of the signal peptide in Escherichia coli. J Biol Chem. 1990;265:8162–8169. - PubMed
-
- Akiyama Y, Kihara A, Tokuda H, Ito K. FtsH (HflB) is an ATP-dependent protease selectively acting on SecY and some other membrane proteins. J Biol Chem. 1996;271:31196–31201. - PubMed
-
- Altamura N, Capitanio N, Bonnefoy N, Papa S, Dujardin G. The Saccharomyces cerevisiae OXA1 gene is required for the correct assembly of cytochrome c oxidase and oligomycin-sensitive ATP synthase. FEBS Lett. 1996;382:111–115. - PubMed
-
- Andersen C L, Matthey-Dupraz A, Missiakas D, Raina S. A new Escherichia coli gene, dsbG, encodes a periplasmic protein involved in disulfide bond formation, required for recycling DsbA/DsbB and DsbC redox proteins. Mol Microbiol. 1997;26:121–132. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

