The human tRNA(m(2)(2)G(26))dimethyltransferase: functional expression and characterization of a cloned hTRM1 gene
- PMID: 10982862
- PMCID: PMC110725
- DOI: 10.1093/nar/28.18.3445
The human tRNA(m(2)(2)G(26))dimethyltransferase: functional expression and characterization of a cloned hTRM1 gene
Abstract
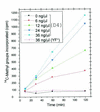
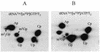
This paper presents the first example of a complete gene sequence coding for and expressing a biologically functional human tRNA methyltransferase: the hTRM1 gene product tRNA(m(2)(2)G)dimethyltransferase. We isolated a human cDNA (1980 bp) made from placental mRNA coding for the full-length (659 amino acids) human TRM1 polypeptide. The sequence was fairly similar to Saccharomyces cerevisiae Trm1p, to Caenorhabditis elegans TRM1p and to open reading frames (ORFs) found in mouse and a plant (Arabidopsis thaliana) DNA. The human TRM1 gene was expressed at low temperature in Escherichia coli as a functional recombinant protein, able to catalyze the formation of dimethylguanosine in E.coli tRNA in vivo. It targeted solely position G(26) in T7 transcribed spliced and unspliced human tRNA(Tyr) in vitro and in yeast trm1 mutant tRNA. Thus, the human TRM1 protein is a tRNA(m(2)(2)G(26))dimethyltransferase. Compared with yeast Trm1p, hTRM1p has a C-terminal protrusion of approximately 90 amino acids which shows similarities to a mouse protein related to RNA splicing. A deletion of these 90 C-terminal amino acids left the modification activity in vitro intact. Among point mutations in the hTRM1 gene, only those located in conserved regions of hTRM1p completely eliminated modification activity.
Figures


References
-
- Björk G.R. (1995) In Söll,D. and Rajbhandary,U.L. (eds), tRNA: Structure, Biosynthesis, and Function. ASM Press, Washington, DC, pp. 165–205.
-
- Johnson G.D., Pirtle,I.L. and Pirtle,R.M. (1985) Arch. Biochem. Biophys., 236, 448–453. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

