Apolipoprotein E variation at the sequence haplotype level: implications for the origin and maintenance of a major human polymorphism
- PMID: 10986041
- PMCID: PMC1287893
- DOI: 10.1086/303070
Apolipoprotein E variation at the sequence haplotype level: implications for the origin and maintenance of a major human polymorphism
Abstract
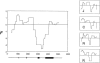
Three common protein isoforms of apolipoprotein E (apoE), encoded by the epsilon2, epsilon3, and epsilon4 alleles of the APOE gene, differ in their association with cardiovascular and Alzheimer's disease risk. To gain a better understanding of the genetic variation underlying this important polymorphism, we identified sequence haplotype variation in 5.5 kb of genomic DNA encompassing the whole of the APOE locus and adjoining flanking regions in 96 individuals from four populations: blacks from Jackson, MS (n=48 chromosomes), Mayans from Campeche, Mexico (n=48), Finns from North Karelia, Finland (n=48), and non-Hispanic whites from Rochester, MN (n=48). In the region sequenced, 23 sites varied (21 single nucleotide polymorphisms, or SNPs, 1 diallelic indel, and 1 multiallelic indel). The 22 diallelic sites defined 31 distinct haplotypes in the sample. The estimate of nucleotide diversity (site-specific heterozygosity) for the locus was 0.0005+/-0.0003. Sequence analysis of the chimpanzee APOE gene showed that it was most closely related to human epsilon4-type haplotypes, differing from the human consensus sequence at 67 synonymous (54 substitutions and 13 indels) and 9 nonsynonymous fixed positions. The evolutionary history of allelic divergence within humans was inferred from the pattern of haplotype relationships. This analysis suggests that haplotypes defining the epsilon3 and epsilon2 alleles are derived from the ancestral epsilon4s and that the epsilon3 group of haplotypes have increased in frequency, relative to epsilon4s, in the past 200,000 years. Substantial heterogeneity exists within all three classes of sequence haplotypes, and there are important interpopulation differences in the sequence variation underlying the protein isoforms that may be relevant to interpreting conflicting reports of phenotypic associations with variation in the common protein isoforms.
Figures
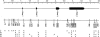




References
Electronic-Database Information
-
- Arlequin, http://lgb.unige.ch/arlequin (for Arlequin program)
-
- DnaSP, http://www.bio.ub.es/~julio/DnaSP.html (for DnaSP software)
-
- Family Blood Pressure Program Web site, http://www.hypertensiongenetics.org
-
- Fluxus Engineering, http://www.fluxus-engineering.com/sharenet.htm (for Network 2.0 software)
-
- Genbank, http://www.ncbi.nlm.nih.gov/entrez/ (for chimpanzee APOE reference sequence [AF261280])
References
-
- Artiga MJ, Bullido MJ, Frank A, Sastre I, Recuero M, Garcia MA, Lendon CL, Han SW, Morris JC, Vazquez J, Goate A, Valdivieso F (1998a) Risk for Alzheimer's disease correlates with transcriptional activity of the APOE gene. Hum Mol Genet 7:1887–1892 - PubMed
-
- Artiga MJ, Bullido MJ, Sastre I, Recuero M, Garcia MA, Aldudo J, Vazquez J, Valdivieso F (1998b) Allelic polymorphisms in the transcriptional regulatory region of apolipoprotein E gene. Fed Europ Bioch Soc Letters 421:105–108 - PubMed
-
- Bullido MJ, Artiga MJ, Recuero M, Sastre I, Garcia MA, Aldudo J, Lendon C, Han SW, Morris JC, Frank A, Vazquez J, Goate A, Valdivieso F (1998) A polymorphism in the regulatory region of APOE associated with risk for Alzheimer's dementia. Nat Genet 18:69–71 - PubMed
-
- Chartier-Harlin MC, Parfitt M, Legrain S, Perez-Tur J, Brousseau T, Evans A, Berr C, Vidal O, Roques P, Gourlet V, Fruchart JC, Delacourte A, Rossor M, Amouyel P (1994) Apolipoprotein E, epsilon 4 allele as a major risk factor for sporadic early and late-onset forms of Alzheimer's disease: analysis of the 19q13.2 chromosomal region. Hum Mol Genet 3:569–574 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

